Introduction
When in April 2019 astronomers first presented the image of a black hole to an astonished world public, it was a pioneering achievement. The radio telescopes used for the recording under the collective name "Event Horizon Telescope" were not new, but they were merely adapted for the project. In the past, the data of several telescopes had already been linked to form a virtual giant telescope. The breakthrough of imaging a black hole for the first time was therefore based more on visions and their implementation with data and computing power than on the use of novel telescopes. With clever algorithms, the ever more powerful computers open up the use of immense amounts of data in order to open up new horizons.
Current developments in microscopy point in a similar direction. Especially in the field of optics and hardware there are groundbreaking innovations: Powerful algorithms open up new ways of handling data and provide additional, valuable information from samples. The combination of different types of data makes it possible to draw relevant conclusions more often and makes complex experiments more practical.
One example of development are the THUNDER Imagers from Leica Microsystems with which scientists can look much deeper into thick samples than was previously possible by combining software and optics using 3D visualization. Based on widefield microscopy, a THUNDER Imager eliminates blurred image information outside the focal plane that clouds the view of thick samples when imaged with camera-based fluorescence microscopes. Previously, the possibility of quantitative evaluation was limited and only possible using more time-consuming methods. During recording, the instrument and software continuously exchange information. The amount of data exchanged far exceeds that which is possible by the input of a human user.
Crystal clear images despite thick samples
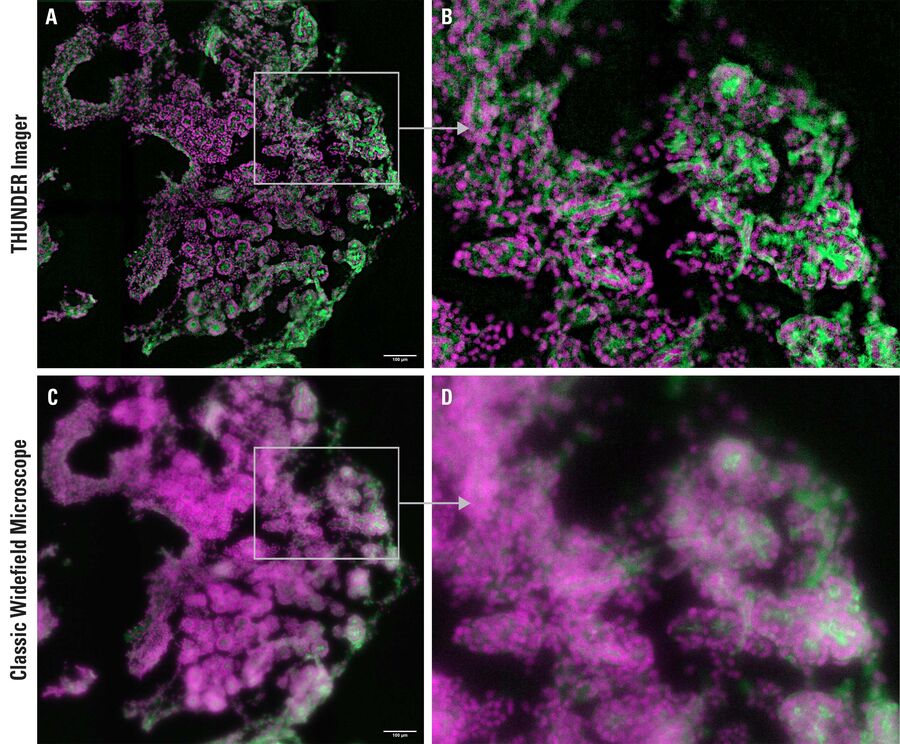
Capturing 3D samples effectively with the speed of camera-based wide field instruments opens up new workflows. An example which shows this advantage is this pancreas study. The focus here is on the self-organisation of individual cells in an organ-like tissue composite, pancreatic implants, and spheroids (Fig. 1).
Clarified embryonic mouse pancreas explants
The 3D cell culture model in combination with mouse genetics and live-cell imaging opens up the possibility of tracking individual cell fate and observing the in vitro development of the organ-like cell culture system. Computational Clearing enables sample-friendly observation and visualization as well as rapid quantitative image analysis of cell differentiation during development.

The resolution of THUNDER imagers is sufficient for many questions in the investigation of spheroids. For such experiments with several positions we are simply faster than with alternative 3D imaging systems".
Dr. Irene Seijo, MPI-CBG Dresden
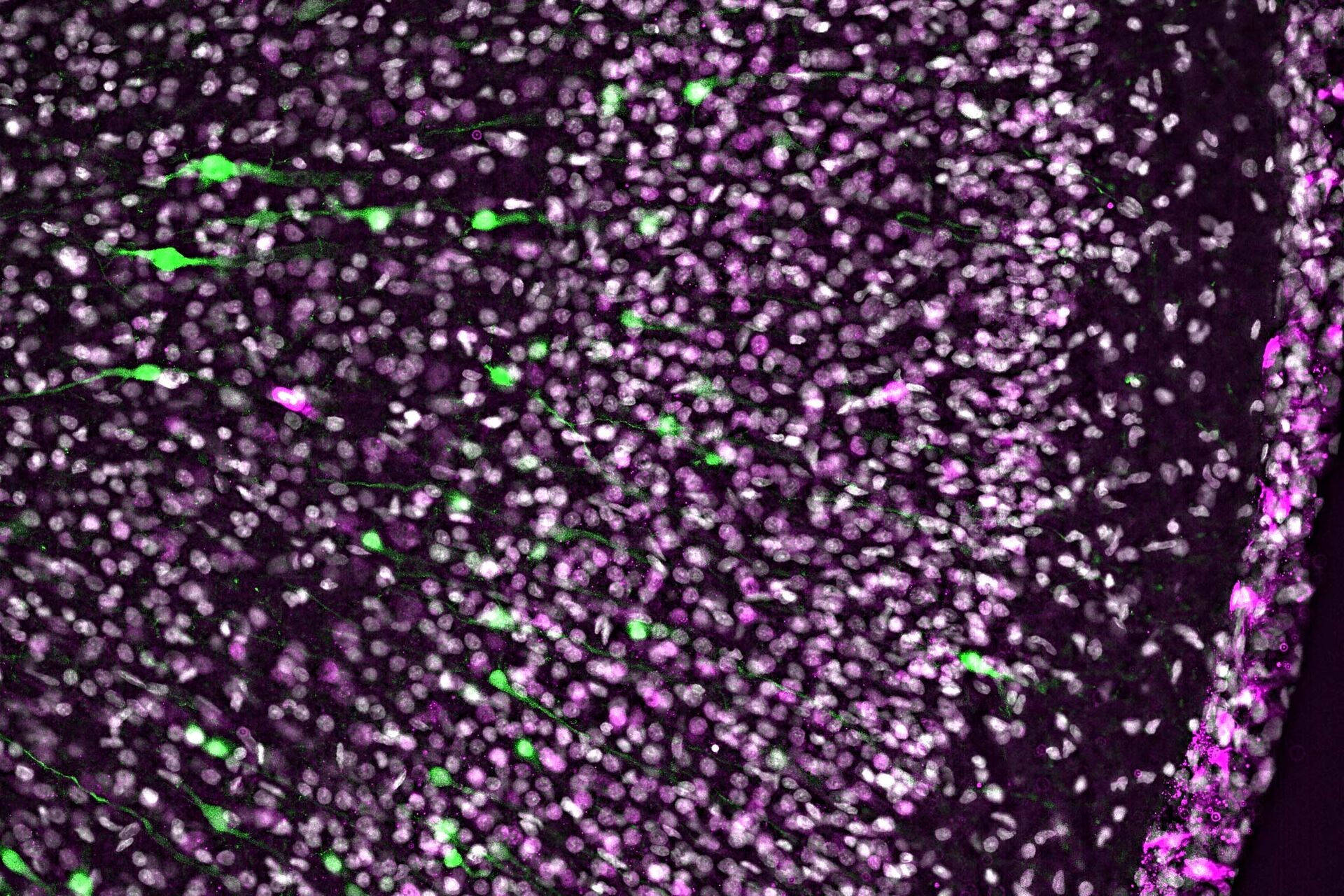
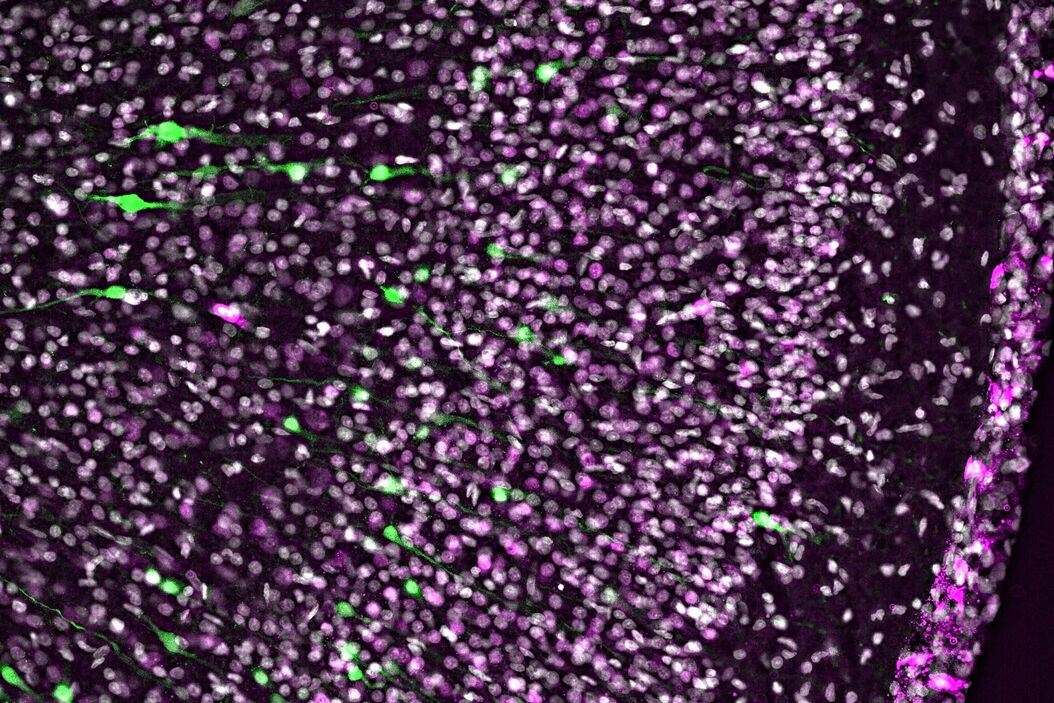
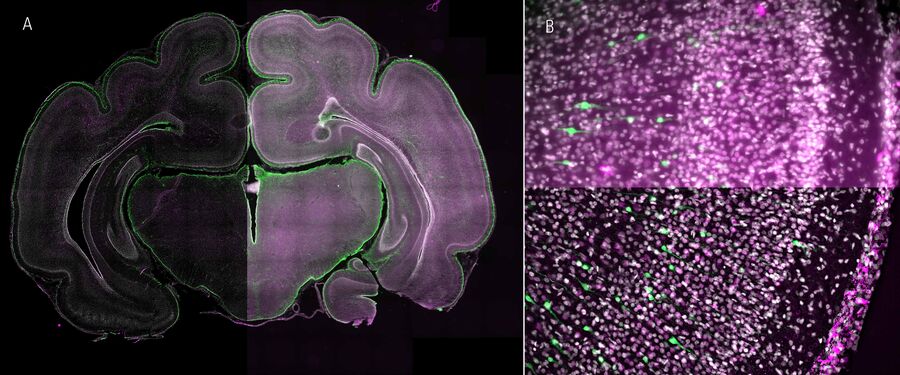
In addition, more and more clarified organs or thick tissue samples are moving into the focus of scientific researchers. For example, certain proteins in ferret neocortex that potentially play a key role in the evolutionary expansion of human cognitive development has been investigated (Fig. 2). Comprehensive screenings are generated using contrast-enhanced, fast-acquired image data from thick tissue samples, saving a great deal of time, and already allow initial evaluation and pre-screening of the data set for further confocal imaging procedures.

With the help of Instant Computational Clearing, I can already localize and evaluate certain signals in brain regions".
Dr. Nereo Kalebic, MPI-CBG Dresden
Developing ferret brain
Data exchange drives science forward
For many scientists it would be advantageous to expand their image data set by gathering images of the same sample from third parties. In this way, evaluations could be carried out on a broader statistical basis in a shorter time or the entire concentration could be directed to the evaluation of existing images. Developing standard file exchange formats for images of biological samples is an important element for consortiums like the OME (Open Microscopy Environment). Only if images from uniformly stored meta information can be automatically found in image databases, for which a comparable evaluation makes sense, can then extensive amounts of data be accessed.
Often large amounts of data are required. If, for example, analyses are to be supported with the help of artificial intelligence, the corresponding systems must first be trained with a large amount of test data for which the result is known. We already see some remarkable examples of the compilation of large amounts of data, such as Emma Lundberg's SciLifeLab in the Human Protein Atlas. The SciLifeLab is a huge database of images that can be analyzed using deep learning techniques to determine patterns in the subcellular localization of proteins.
Collaboration supports protein research
Leica Microsystems recently sponsored a competition on the Kaggle platform that promotes the exchange of data-science know-how. The aim of the competition was to develop an algorithm that sorts the images of the Human Protein Atlas into 28 classes, each showing different organelles of the cell. There were 2,169 teams that faced the challenge of predicting where a protein had migrated to in the cell - with only one image as a starting point. Such a task is particularly difficult, because some proteins go to several locations at the same time and some patterns occur very rarely. In order to be successful, the data scientists had to search for independent data available on the internet in order to supplement the data from the competition. Such efforts help to understand the role of proteins in health and disease at the cellular level.
Conclusion
Although new ways of science can be explored with new devices, data, and software are playing an increasingly important role alongside this almost classical scheme and are introducing something like a "quiet" revolution. Innovators are needed who contribute to the evolution of current methods and go beyond existing horizons with new approaches. It is necessary to supplement powerful tools with which microscopy can discover new horizons for our understanding in the future.
Related Articles
-
Overcoming Observational Challenges in Organoid 3D Cell Culture
Learn how to overcome challenges in observing organoid growth. Read this article and discover new…
Apr 08, 2024Read article -
How do Cells Talk to Each Other During Neurodevelopment?
Professor Silvia Capello presents her group’s research on cellular crosstalk in neurodevelopmental…
Apr 03, 2024Read article -
How to Get Deeper Insights into your Organoid and Spheroid Models
In this eBook, learn about key considerations for imaging 3D cultures, such as organoids and…
Nov 22, 2023Read article