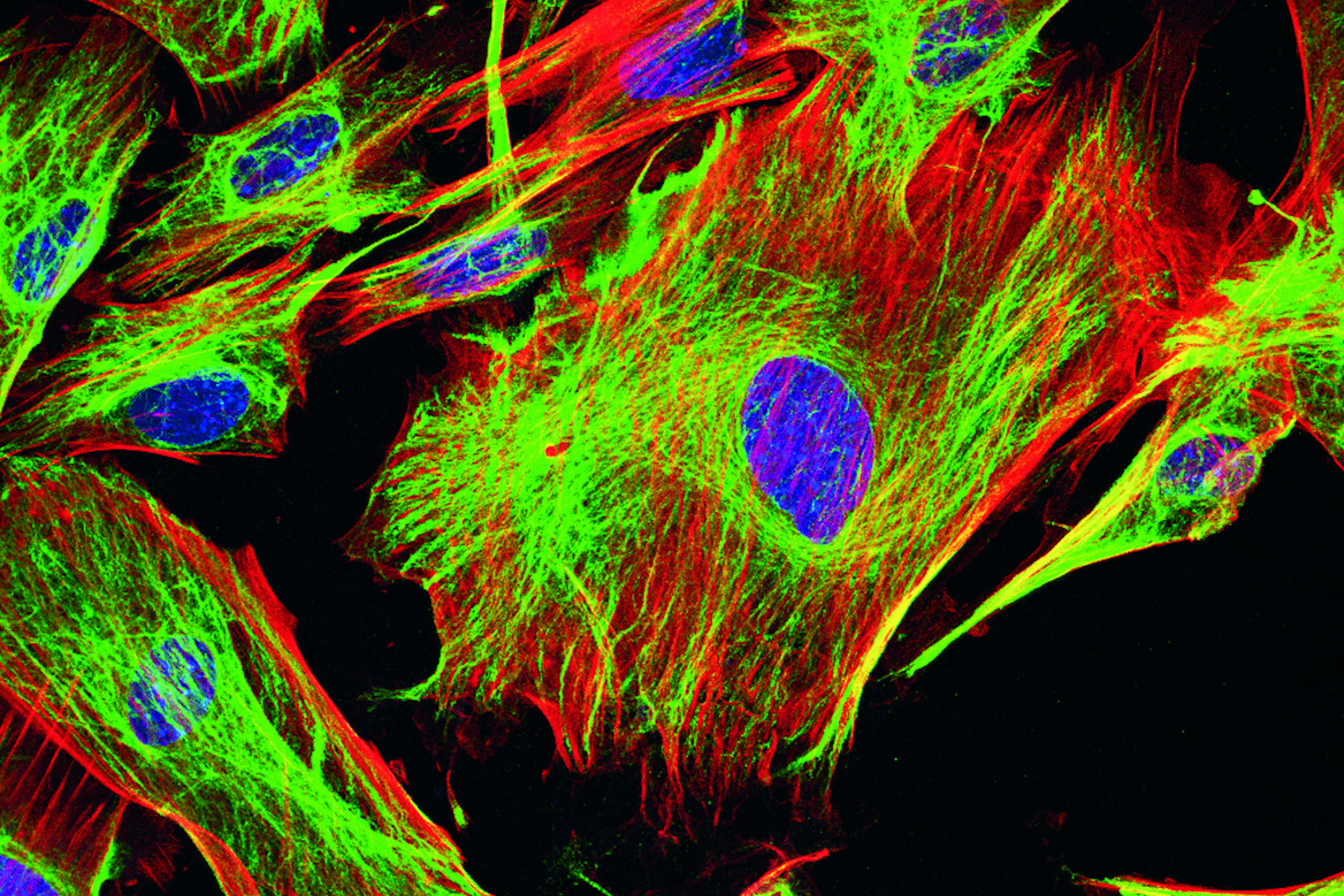
Immunofluorescence
In fluorescence microscopy there are two ways to visualize your protein of interest. Either with the help of an intrinsic fluorescent signal - by genetically linking a fluorescent protein to a target protein - or with the help of fluorescently labeled antibodies that bind specifically to a protein of interest. For some biological questions it is more useful or even necessary to perform the latter one. In the case of histological samples, for example, it is not possible to use fluorescent proteins because in general the specimen is derived from an organism which does not hold any fluorescent proteins. Furthermore, if a functioning antibody is available, immunofluorescence is much faster than fluorescent protein techniques, where you have to clone the gene of interest and transfect DNA into the adequate cell. Another disadvantage of fluorescent proteins lies in their nature of being a protein themselves. With it, they have specific proteinous characteristics inside a cell which can lead to dysfunction or misinterpretations concerning the attached protein of interest. However, it should be considered that using fluorescent proteins is generally the method of choice to study living cells.
Immunofluorescence makes use of the very specific binding affinity of an antibody to its antigen. This can have two different appearances. The easiest way is to use one fluorescently labeled antibody which is binding to the protein of interest. This is called direct immunofluorescence.
In most cases there are two forms of antibodies used. The first one binds to the target protein and is not fluorescently labeled itself (primary antibody). But a second one (secondary antibody) which is binding the primary antibody specifically carries a fluorescent dye. This method is called indirect immunofluorescence and has several advantages. On the one hand there is an amplifying effect, because more than one secondary antibody binds to one primary antibody. On the other hand, it is generally easier to find secondary antibodies than the specific primary antibody labeled with a common fluorescent dye. Below, we review the most commonly used fluorescent dyes.
FITC and TRITC
Fluorescein isothiocyanate (FITC) is an organic fluorescent dye and probably one of the most commonly used in immunofluorescence and flow cytometry. It has an excitation/emission peak at 495/517 nm and can be coupled to distinct antibodies with the help of its reactive isothiocyanate group, which is binding to amino, sulfhydryl, imidazoyl, tyrosyl or carbonyl groups on proteins. FITC (was one of the first dyes which was used for fluorescence microscopy and served as a precursor for other fluorescent dyes like Alexa Fluor®488. Its fluorescence activity is due to its large conjugated aromatic electron system, which is excited by light in the blue spectrum.
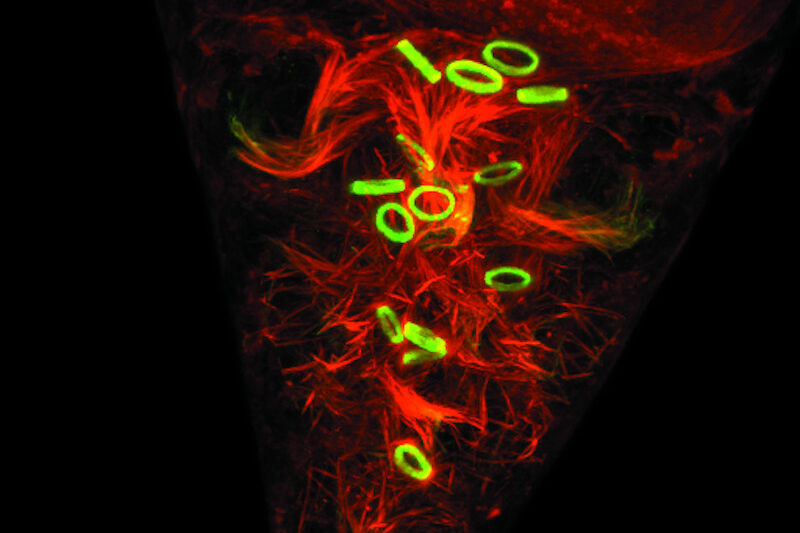
A dye very often used in combination with FITC is TRITC (Tetramethylrhodamine-5-(and 6)-isothiocyanate). In contrast to FITC, TRITC is not a fluorescein but a derivate of the Rhodamine family. Rhodamines also have a large conjugated aromatic electron system, what leads to their fluorescent behavior. TRITC is excited with light in the green spectrum with a maximum at 550 nm. Its emission maximum is lying at 573 nm. The bond to proteins (e.g. antibodies) is also based on a reactive isothiocyanate group.
Even though FITC and TRITC are still widely used, they are rather weak fluorescent dyes and not recommended for state-of-the-art microscopy.
Cyanines
Alexa Fluor® dyes are a big group of negatively charged and hydrophilic fluorescent dyes, frequently used in fluorescence microscopy. All the Alexa Fluor® dyes are sulfonated forms of different basic fluorescent substances like fluorescein, coumarin, cyanine or rhodamine (e.g. Alexa Fluor®546, Alexa Fluor®633). The respective laser excitation wavelength is mentioned in their labeling. For example, Alexa Fluor®488, one of the most commonly used dyes, has an excitation maximum at 493 nm, which allows excitation with a standard 488 nm laser, and an emission maximum at 519 nm Alexa Fluor®488 is a fluorescein derivate and has similar properties than FITC. However, it shows better stability, brightness and lower pH sensitivity.
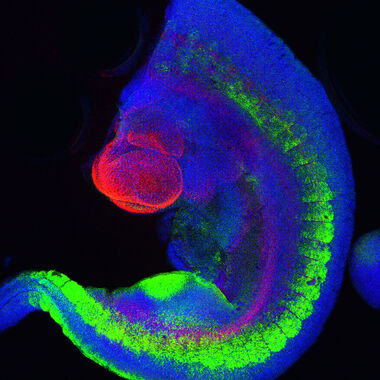
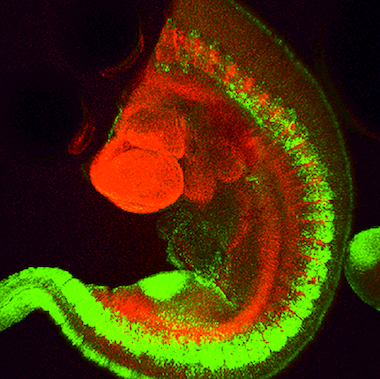
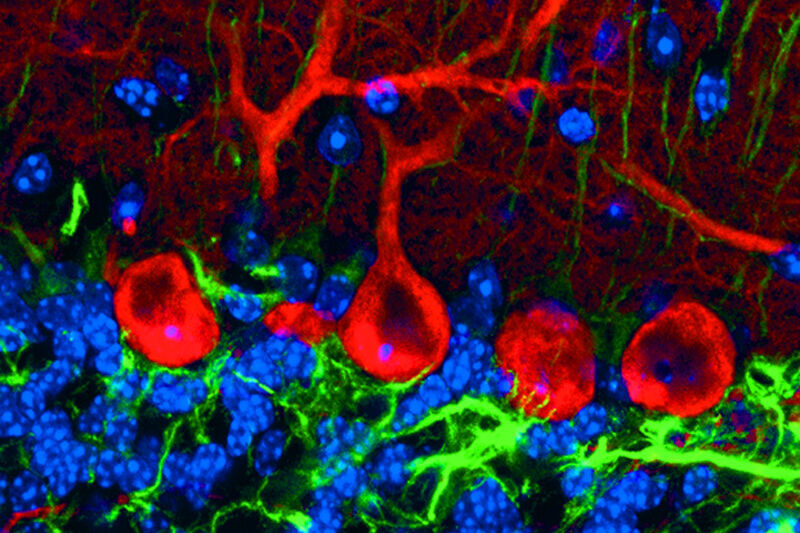
Fig. 2: Mouse transgenic embryo, interlimb somites, Five interlimb somites of an E10.5 mouse transgenic embryo: EpaxialMyf5 eGFP; immuno-stained for GFP-Alexa 488; embryonic muscle fibers stained with Desmin-Cy3, the nuclei are revealed with Hoechst Size from top to bottom: 3.5 mm (a), 800 µm (b). Courtesy of: Aurélie Jory, Cellules Souches et Développement, Institut Pasteur, Paris, France and Imaging centre of IGBMC, IGBMC
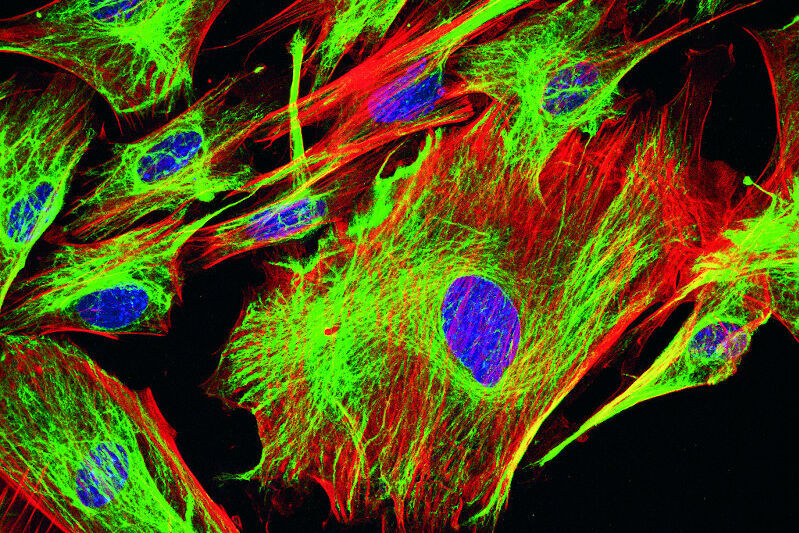
DNA staining
You might want to study nucleic acids using fluorescent microscopy. For example, to define the exact position or number of cells by the detection of their nucleus. One of the most common DNA stains is DAPI (4',6-diamidino-2-phenylindole) which binds to A-T rich regions of the DNA double helix. DAPI fluorescence intensity increases if attached to DNA compared to its unbound state. It is excited by UV-light with a maximum at 358 nm. Emission spectrum is broad and peaks at 461 nm. A weak fluorescence can also be detected for RNA binding. In this case, emission shifts to 500 nm. Interestingly, DAPI is able to permeate an intact plasma membrane which makes it useful for fixed and living cells.
A second broadly used DNA stain option is the family of Hoechst dyes. Hoechst 33258, Hoechst 33342, and Hoechst 34580 are Bisbenzimides with intercalating tendency to A-T rich areas. Similar to DAPI they are excited by UV-light and have an emission maximum at 455 nm which is shifted to 510–540 nm in an unbound condition. Hoechst stains are also cell permeable and can therefore be used in fixed and living cells. They exhibit lower toxicity than DAPI.
A membrane-impermeable DNA stain is Propidium-Iodide which is often used to differentiate between living and dead cells in a cell culture because it cannot enter an intact cell. Propidium-Iodide is also an intercalating agent but with no binding preference for distinct bases. In the nucleic acid bound state, its excitation maximum is at 538 nm. Highest emission is at 617 nm. Unbound Propidium-Iodide excitation and emission maxima are shifted to lower wavelengths and lower intensity. It can also bind to RNA without changing its fluorescent characteristics. To distinguish DNA from RNA it is necessary to use adequate nucleases.
A dye which is capable to make a difference between DNA and RNA without previous manipulation is Acridine Orange. Its excitation/emission maximum pair is 502 nm/525 nm in the DNA bound version and turns to 460 nm/650 nm in the RNA bound state. Furthermore, it can enter acidic compartments like lysosomes where the cationic dye is protonated. In this acidic surrounding Acridine Orange is excited by light in the blue spectrum, whereas emission is strongest in the orange region. It is often used to identify apoptotic cells, as they have a lot of engulfed acidic compartments
Compartment and organelle specific dyes
There is a number of specific dyes to study cell compartments such as lysosomes, endosomes or organelles such as mitochondria.
One well known way to observe mitochondria is the utilization of MitoTracker®. This is a cell permeable dye with a mildly thiol-reactive chloromethyl moiety used to bind to matrix proteins covalently by reacting with free thiol groups of cysteine residues. MitoTracker® exists in different colours and modifications (s. Table 1) . In contrast to conventional mitochondria specific stains like rhodamine 123 or tetramethylrosamine, MitoTracker® is not washed out after destruction of the membrane potential with fixatives.
LysoTracker is a group of dyes available in different colors used to stain acidic compartments such as lysosomes. These are membrane permeable weak bases linked to a fluorophore. Most probably these bases have an affinity to acidic compartments because of protonation (s. Table 1).
A comparable compartment to lysosomes is the vacuole in fungi like Saccharomyces cerevisiae. This membrane enclosed space is also of an acidic nature. One way to visualize it in fluorescence microscopy is the use of styryl-based dyes like FM 4-64® or FM 5-95®.
The Endoplasmic Reticulum (ER) is usually stained when studying protein secretion. One classical dye to stain this compartment is DiOC6(3) which has a preference for the ER but still binds to other membranes like those of mitochondria. Another way to specifically stain the ER is to use ER-Trackers like ER-Tracker Green and Red. Both are BODIPY based dyes which are linked to glibenclamide – a sulfonylurease – which binds to ATP sensitive Potassium channels exclusively resident in the ER membrane. BODIPY (boron-dipyrromethene) describes a group of relatively pH insensitive dyes which are almost all water insoluble. This makes them a very good tool for lipid and membrane labeling.
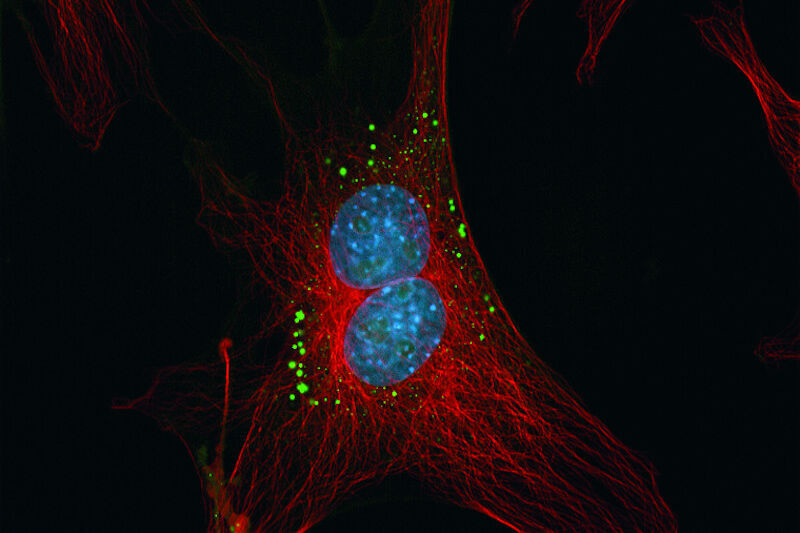
The adjacent compartment to the ER – the Golgi apparatus – can be labeled with fluorescent ceramide analogs like NBD C6-ceramide and BODIPY FL C5-ceramide. Ceramides are Sphingolipids which are highly enriched in the Golgi apparatus.
With the help of further lipid-based dyes it is possible to stain special membrane regions like lipid-rafts. These cholesterol rich domains can be visualized by using NBD-6 Cholestrol or NBP-12 Cholesterol amongst others (Avanti Polar Lipids).
It is also possible to stain the area of interest with the help of proteins with preferences for distinct locations in the cell. One example is the use of wheat germ agglutinin (WGA) which binds specifically to sialic acid and N-acetylglucosaminyl present in the plasma membrane.
Ion imaging
In the case of neuronal studies, gene activity or cellular movement it is of interest to study the ion concentration of the cell. Sodium, calcium, chloride or magnesium ions have a deep impact on many different cellular events. Typically, ions can be trapped with the help of fluorescently labeled chelators that change their spectral properties when bound to the appropriate ions. One example of labelled chelators are the calcium indicators fura-2, indo-1, fluo-3, fluo-4 and Calcium-Green. For sodium detection, SBFI (sodium-binding benzofurzanisophthalate) or Sodium Green are commonly used. PBFI (potassium-binding benzofurzanisophthalate) detects potassium ions.
Functional assays
“Functional assays” is the broad term used to cover standardized experiments to assess various functions that can be visualized with fluorescent markers. These markers can encompass but is not limited to any of the above-mentioned labeling techniques and fluorophores. For many of these functional assays, staining kits are commercially available and can easily be applied to a vast variety of samples. One example for a functional assay is the commonly known and widely used live dead assay. Two fluorophores are utilized to label live cells and dead cells. Having both values at hand the overall health status of the cells can be assessed. Correlating this information with additional markers may even increase the understanding of the underly process.
Fluorescent dyes and their excitation and emission wavelength peaks
The table below shows a comprehensive list of fluorescent dyes with their respective excitation and emission wavelength peaks. Please note that besides those peaks, every dye features distinct excitation and emission spectra. When selecting several dyes to use in combination in one experiment, researchers should be aware of overlapping excitation and emission spectra due to crosstalk (or bleedthrough), which can result in false negatives or positives, or otherwise obscure data. Another source that can distort fluorescence imaging is autofluorescence by naturally occurring fluorescent proteins in cells and tissues, which particularly needs to be considered in experiments with plants or algae. Good understanding of the spectra of the dyes used in the sample is also important to choose the right light source for excitation (e.g. LED, arc lamps, laser lines) and the right filters and detectors for emission.
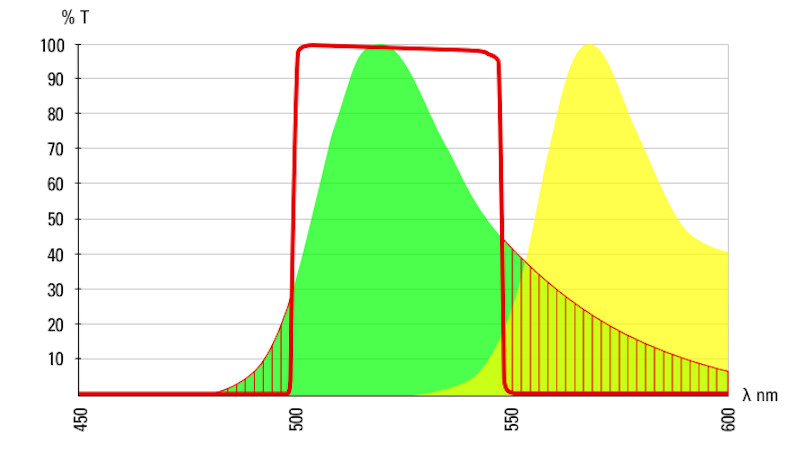
indicates the band pass of a 488 emissions filter.
Table 1
| Sample Fluorescent Dyes | Excitation | Emission |
| Indo-1, Ca saturated | 331 nm | 404 nm |
| Indo-1 Ca2+ | 346 nm | 404 nm |
| Cascade Blue BSA pH 7.0 | 401 nm | 419 nm |
| Cascade Blue | 398 nm | 420 nm |
| LysoTracker Blue | 373 nm | 421 nm |
| Alexa 405 | 401 nm | 421 nm |
| LysoSensor Blue pH 5.0 | 374 nm | 424 nm |
| LysoSensor Blue | 374 nm | 424 nm |
| DyLight 405 | 399 nm | 434 nm |
| DyLight 350 | 332 nm | 435 nm |
| BFP (Blue Fluorescent Protein) | 380 nm | 439 nm |
| Alexa 350 | 343 nm | 441 nm |
| 7-Amino-4-methylcoumarin pH 7.0 | 346 nm | 442 nm |
| Amino Coumarin | 345 nm | 442 nm |
| AMCA conjugate | 347 nm | 444 nm |
| Coumarin | 360 nm | 447 nm |
| 7-Hydroxy-4-methylcoumarin | 360 nm | 447 nm |
| 7-Hydroxy-4-methylcoumarin pH 9.0 | 361 nm | 448 nm |
| 6,8-Difluoro-7-hydroxy-4-methylcoumarin pH 9.0 | 358 nm | 450 nm |
| Hoechst 33342 | 352 nm | 455 nm |
| Pacific Blue | 404 nm | 455 nm |
| Hoechst 33258 | 352 nm | 455 nm |
| Hoechst 33258-DNA | 352 nm | 455 nm |
| Pacific Blue antibody conjugate pH 8.0 | 404 nm | 455 nm |
| PO-PRO-1 | 434 nm | 457 nm |
| PO-PRO-1-DNA | 435 nm | 457 nm |
| POPO-1 | 433 nm | 457 nm |
| POPO-1-DNA | 433 nm | 458 nm |
| DAPI-DNA | 359 nm | 461 nm |
| DAPI | 358 nm | 463 nm |
| Marina Blue | 362 nm | 464 nm |
| SYTOX Blue-DNA | 445 nm | 470 nm |
| CFP (Cyan Fluorescent Protein) | 434 nm | 474 nm |
| eCFP (Enhanced Cyan Fluorescent Protein) | 437 nm | 476 nm |
| 1-Anilinonaphthalene-8-sulfonic acid (1,8-ANS) | 375 nm | 479 nm |
| Indo-1, Ca free | 346 nm | 479 nm |
| 1,8-ANS (1-Anilinonaphthalene-8-sulfonic acid) | 375 nm | 480 nm |
| BO-PRO-1-DNA | 462 nm | 482 nm |
| BOPRO-1 | 462 nm | 482 nm |
| BOBO-1-DNA | 461 nm | 484 nm |
| SYTO 45-DNA | 451 nm | 486 nm |
| evoglow-Pp1 | 448 nm | 495 nm |
| evoglow-Bs1 | 448 nm | 496 nm |
| evoglow-Bs2 | 448 nm | 496 nm |
| Auramine O | 431 nm | 501 nm |
| DiO | 487 nm | 501 nm |
| LysoSensor Green pH 5.0 | 447 nm | 502 nm |
| Cy 2 | 489 nm | 503 nm |
| LysoSensor Green | 447 nm | 504 nm |
| Fura-2, high Ca | 336 nm | 504 nm |
| Fura-2 Ca2+sup> | 336 nm | 505 nm |
| SYTO 13-DNA | 488 nm | 506 nm |
| YO-PRO-1-DNA | 491 nm | 507 nm |
| YOYO-1-DNA | 491 nm | 509 nm |
| eGFP (Enhanced Green Fluorescent Protein) | 488 nm | 509 nm |
| LysoTracker Green | 503 nm | 509 nm |
| GFP (S65T) | 489 nm | 509 nm |
| BODIPY FL, MeOH | 502 nm | 511 nm |
| Sapphire | 396 nm | 511 nm |
| BODIPY FL conjugate | 503 nm | 512 nm |
| MitoTracker Green | 490 nm | 512 nm |
| MitoTracker Green FM, MeOH | 490 nm | 512 nm |
| Fluorescein 0.1 M NaOH | 493 nm | 513 nm |
| Calcein pH 9.0 | 494 nm | 514 nm |
| Fluorescein pH 9.0 | 490 nm | 514 nm |
| Calcein | 493 nm | 514 nm |
| Fura-2, no Ca | 367 nm | 515 nm |
| Fluo-4 | 494 nm | 516 nm |
| FDA | 495 nm | 517 nm |
| DTAF | 495 nm | 517 nm |
| Fluorescein | 495 nm | 517 nm |
| Fluorescein antibody conjugate pH 8.0 | 493 nm | 517 nm |
| CFDA | 495 nm | 517 nm |
| FITC | 495 nm | 517 nm |
| Alexa Fluor 488 hydrazide-water | 493 nm | 518 nm |
| DyLight 488 | 493 nm | 518 nm |
| 5-FAM pH 9.0 | 492 nm | 518 nm |
| FITC antibody conjugate pH 8.0 | 495 nm | 519 nm |
| Alexa 488 | 493 nm | 520 nm |
| Rhodamine 110 | 497 nm | 520 nm |
| Rhodamine 110 pH 7.0 | 497 nm | 520 nm |
| Acridine Orange | 431 nm | 520 nm |
| Alexa Fluor 488 antibody conjugate pH 8.0 | 499 nm | 520 nm |
| BCECF pH 5.5 | 485 nm | 521 nm |
| PicoGreendsDNA quantitation reagent | 502 nm | 522 nm |
| SYBR Green I | 498 nm | 522 nm |
| Rhodaminen Green pH 7.0 | 497 nm | 523 nm |
| CyQUANT GR-DNA | 502 nm | 523 nm |
| NeuroTrace 500/525, green fluorescent Nissl stain-RNA | 497 nm | 524 nm |
| DansylCadaverine | 335 nm | 524 nm |
| Rhodol Green antibody conjugate pH 8.0 | 499 nm | 524 nm |
| Fluoro-Emerald | 495 nm | 524 nm |
| Nissl | 497 nm | 524 nm |
| Fluorescein dextran pH 8.0 | 501 nm | 524 nm |
| Rhodamine Green | 497 nm | 524 nm |
| 5-(and-6)-Carboxy-2', 7'-dichlorofluorescein pH 9.0 | 504 nm | 525 nm |
| DansylCadaverine, MeOH | 335 nm | 526 nm |
| eYFP (Enhanced Yellow Fluorescent Protein) | 514 nm | 526 nm |
| Oregon Green 488 | 498 nm | 526 nm |
| Oregon Green 488 antibody conjugate pH 8.0 | 498 nm | 526 nm |
| Fluo-3 | 506 nm | 527 nm |
| BCECF pH 9.0 | 501 nm | 527 nm |
| SBFI-Na+ | 336 nm | 527 nm |
| Fluo-3 Ca2+ | 506 nm | 527 nm |
| Rhodamine 123, MeOH | 507 nm | 529 nm |
| FlAsH | 509 nm | 529 nm |
| Calcium Green-1 Ca2+ | 506 nm | 529 nm |
| Magnesium Green | 507 nm | 530 nm |
| DM-NERF pH 4.0 | 493 nm | 530 nm |
| Calcium Green | 506 nm | 530 nm |
| Citrine | 515 nm | 530 nm |
| LysoSensor Yellow pH 9.0 | 335 nm | 530 nm |
| TO-PRO-1-DNA | 515 nm | 531 nm |
| Magnesium Green Mg2+ | 507 nm | 531 nm |
| Sodium Green Na+ | 507 nm | 531 nm |
| TOTO-1-DNA | 514 nm | 531 nm |
| Oregon Green 514 | 512 nm | 532 nm |
| Oregon Green 514 antibody conjugate pH 8.0 | 513 nm | 533 nm |
| NBD-X | 466 nm | 534 nm |
| DM-NERF pH 7.0 | 509 nm | 537 nm |
| NBD-X, MeOH | 467 nm | 538 nm |
| CI-NERF pH 6.0 | 513 nm | 538 nm |
| Alexa 430 | 431 nm | 540 nm |
| Alexa Fluor 430 antibody conjugate pH 7.2 | 431 nm | 540 nm |
| CI-NERF pH 2.5 | 504 nm | 541 nm |
| Lucifer Yellow, CH | 428 nm | 542 nm |
| LysoSensor Yellow pH 3.0 | 389 nm | 542 nm |
| 6-TET, SE pH 9.0 | 521 nm | 542 nm |
| Eosin antibody conjugate pH 8.0 | 525 nm | 546 nm |
| Eosin | 524 nm | 546 nm |
| 6-Carboxyrhodamine 6G pH 7.0 | 526 nm | 547 nm |
| 6-Carboxyrhodamine 6G, hydrochloride | 525 nm | 547 nm |
| Bodipy R6G SE | 528 nm | 547 nm |
| BODIPY R6G, MeOH | 528 nm | 547 nm |
| 6 JOE | 520 nm | 548 nm |
| Cascade Yellow antibody conjugate pH 8.0 | 399 nm | 549 nm |
| Cascade Yellow | 399 nm | 549 nm |
| mBanana | 540 nm | 553 nm |
| Alexa Fluor 532 antibody conjugate pH 7.2 | 528 nm | 553 nm |
| Alexa 532 | 528 nm | 553 nm |
| Erythrosin-5-isothiocyanate pH 9.0 | 533 nm | 554 nm |
| 6-HEX, SE pH 9.0 | 534 nm | 559 nm |
| mOrange | 548 nm | 562 nm |
| mHoneydew | 478 nm | 562 nm |
| Cy 3 | 549 nm | 562 nm |
| Rhodamine B | 543 nm | 565 nm |
| DiI | 551 nm | 565 nm |
| 5-TAMRA-MeOH | 543 nm | 567 nm |
| Alexa 555 | 553 nm | 568 nm |
| Alexa Fluor 555 antibody conjugate pH 7.2 | 553 nm | 568 nm |
| DyLight 549 | 555 nm | 569 nm |
| BODIPY TMR-X, SE | 544 nm | 570 nm |
| BODIPY TMR-X, MeOH | 544 nm | 570 nm |
| PO-PRO-3-DNA | 539 nm | 571 nm |
| PO-PRO-3 | 539 nm | 571 nm |
| Rhodamine | 551 nm | 573 nm |
| Bodipy TMR-X conjugate | 544 nm | 573 nm |
| POPO-3 | 533 nm | 573 nm |
| Alexa 546 | 562 nm | 573 nm |
| BODIPY TMR-X antibody conjugate pH 7.2 | 544 nm | 573 nm |
| Calcium Orange Ca2+ | 549 nm | 573 nm |
| TRITC | 550 nm | 573 nm |
| Calcium Orange | 549 nm | 574 nm |
| Rhodaminephalloidin pH 7.0 | 558 nm | 575 nm |
| MitoTracker Orange | 551 nm | 575 nm |
| MitoTracker Orange, MeOH | 551 nm | 575 nm |
| Phycoerythrin | 565 nm | 575 nm |
| Magnesium Orange | 550 nm | 575 nm |
| R-Phycoerythrin pH 7.5 | 565 nm | 576 nm |
| 5-TAMRA pH 7.0 | 553 nm | 576 nm |
| 5-TAMRA | 549 nm | 577 nm |
| Rhod-2 | 552 nm | 577 nm |
| FM 1-43 | 472 nm | 578 nm |
| Rhod-2 Ca2+ | 553 nm | 578 nm |
| Tetramethylrhodamine antibody conjugate pH 8.0 | 552 nm | 578 nm |
| FM 1-43 lipid | 473 nm | 579 nm |
| LOLO-1-DNA | 568 nm | 580 nm |
| dTomato | 554 nm | 581 nm |
| DsRed | 563 nm | 581 nm |
| Dapoxyl (2-aminoethyl) sulfonamide | 372 nm | 582 nm |
| Tetramethylrhodamine dextran pH 7.0 | 555 nm | 582 nm |
| Fluor-Ruby | 554 nm | 582 nm |
| Resorufin | 571 nm | 584 nm |
| Resorufin pH 9.0 | 571 nm | 584 nm |
| mTangerine | 568 nm | 585 nm |
| LysoTracker Red | 578 nm | 589 nm |
| Lissaminerhodamine | 572 nm | 590 nm |
| Cy 3.5 | 578 nm | 591 nm |
| Rhodamine Red-X antibody conjugate pH 8.0 | 573 nm | 591 nm |
| Sulforhodamine 101, EtOH | 578 nm | 593 nm |
| JC-1 pH 8.2 | 593 nm | 595 nm |
| JC-1 | 592 nm | 595 nm |
| mStrawberry | 575 nm | 596 nm |
| MitoTracker Red | 578 nm | 599 nm |
| MitoTracker Red, MeOH | 578 nm | 599 nm |
| X-Rhod-1 Ca2+ | 580 nm | 602 nm |
| Alexa Fluor 568 antibody conjugate pH 7.2 | 579 nm | 603 nm |
| Alexa 568 | 576 nm | 603 nm |
| 5-ROX pH 7.0 | 578 nm | 604 nm |
| 5-ROX (5-Carboxy-X-rhodamine, triethylammonium salt) | 578 nm | 604 nm |
| BO-PRO-3-DNA | 574 nm | 604 nm |
| BOPRO-3 | 574 nm | 604 nm |
| BOBO-3-DNA | 570 nm | 605 nm |
| Ethidium Bromide | 524 nm | 605 nm |
| ReAsH | 597 nm | 608 nm |
| Calcium Crimson | 589 nm | 608 nm |
| Calcium Crimson Ca2+ | 590 nm | 608 nm |
| mRFP | 585 nm | 608 nm |
| mCherry | 587 nm | 610 nm |
| Texas Red-X antibody conjugate pH 7.2 | 596 nm | 613 nm |
| HcRed | 590 nm | 614 nm |
| DyLight 594 | 592 nm | 616 nm |
| Ethidium homodimer-1-DNA | 528 nm | 617 nm |
| Ethidiumhomodimer | 528 nm | 617 nm |
| Propidium Iodide | 538 nm | 617 nm |
| SYPRO Ruby | 467 nm | 618 nm |
| Propidium Iodide-DNA | 538 nm | 619 nm |
| Alexa 594 | 590 nm | 619 nm |
| BODIPY TR-X, SE | 588 nm | 621 nm |
| BODIPY TR-X, MeOH | 588 nm | 621 nm |
| BODIPY TR-X phallacidin pH 7.0 | 590 nm | 621 nm |
| Alexa Fluor 610 R-phycoerythrin streptavidin pH 7.2 | 567 nm | 627 nm |
| YO-PRO-3-DNA | 613 nm | 629 nm |
| Di-8 ANEPPS | 469 nm | 630 nm |
| Di-8-ANEPPS-lipid | 469 nm | 631 nm |
| YOYO-3-DNA | 612 nm | 631 nm |
| Nile Red-lipid | 553 nm | 636 nm |
| Nile Red | 559 nm | 637 nm |
| DyLight 633 | 624 nm | 646 nm |
| mPlum | 587 nm | 649 nm |
| TO-PRO-3-DNA | 642 nm | 657 nm |
| DDAO pH 9.0 | 648 nm | 657 nm |
| Fura Red, high Ca | 434 nm | 659 nm |
| Allophycocyanin pH 7.5 | 651 nm | 660 nm |
| APC (allophycocyanin) | 650 nm | 660 nm |
| Nile Blue, EtOH | 631 nm | 660 nm |
| TOTO-3-DNA | 642 nm | 661 nm |
| Cy 5 | 646 nm | 664 nm |
| BODIPY 650/665-X, MeOH | 646 nm | 664 nm |
| Alexa Fluor 647 R-phycoerythrin streptavidin pH 7.2 | 569 nm | 666 nm |
| DyLight 649 | 652 nm | 668 nm |
| Alexa Fluor 647 antibody conjugate pH 7.2 | 653 nm | 668 nm |
| Alexa 647 | 653 nm | 669 nm |
| Fura Red Ca2+ | 435 nm | 670 nm |
| Atto 647 | 644 nm | 670 nm |
| Fura Red, low Ca | 472 nm | 673 nm |
| Carboxynaphthofluorescein pH 10.0 | 600 nm | 674 nm |
| Alexa 660 | 664 nm | 691 nm |
| Alexa Fluor 660 antibody conjugate pH 7.2 | 663 nm | 691 nm |
| Cy 5.5 | 673 nm | 692 nm |
| Alexa Fluor 680 antibody conjugate pH 7.2 | 679 nm | 702 nm |
| Alexa 680 | 679 nm | 703 nm |
| DyLight 680 | 678 nm | 706 nm |
| Alexa Fluor 700 antibody conjugate pH 7.2 | 696 nm | 719 nm |
| Alexa 700 | 696 nm | 720 nm |
| FM 4-64, 2% CHAPS | 506 nm | 751 nm |
| FM 4-64 | 508 nm | 751 nm |
Related Articles
-
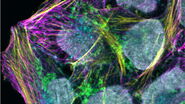
A New World of Confocal Applications with the Next Generation White Light Lasers
As biological questions get more complex, there is an increasing need to study multiple events…
Nov 19, 2020Read article -
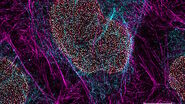
The Guide to STED Sample Preparation
This guide is intended to help users optimize sample preparation for stimulated emission depletion…
Mar 05, 2024Read article -
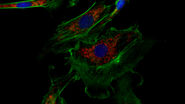
Studying Virus Replication with Fluorescence Microscopy
The results from research on SARS-CoV-2 virus replication kinetics, adaption capabilities, and…
Nov 15, 2023Read article
Related Pages
-

Compound Light Microscopes
Compound light microscopes from Leica Microsystems meet the highest demands whatever the application…
Visit related page -
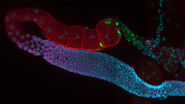
Fluorescence
Find out how fluorescence microscopes from Leica Microsystems support your research. Fluorescence is…
Visit related page