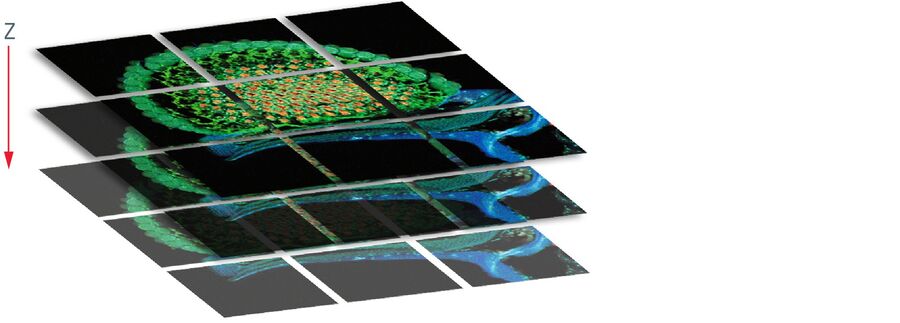
Large specimen albeit small scanning area
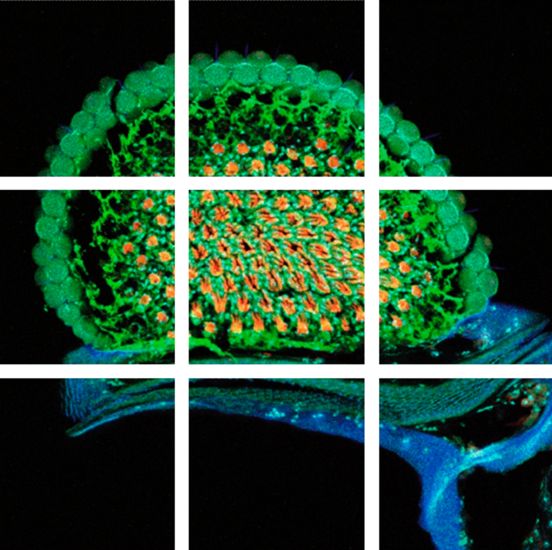
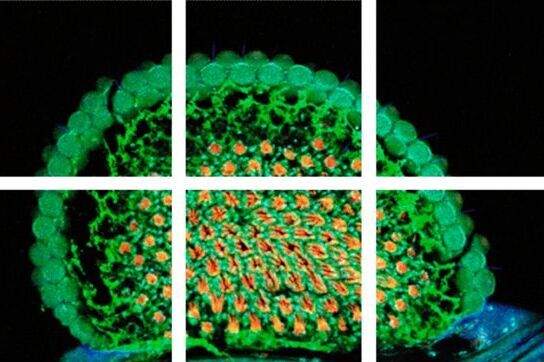
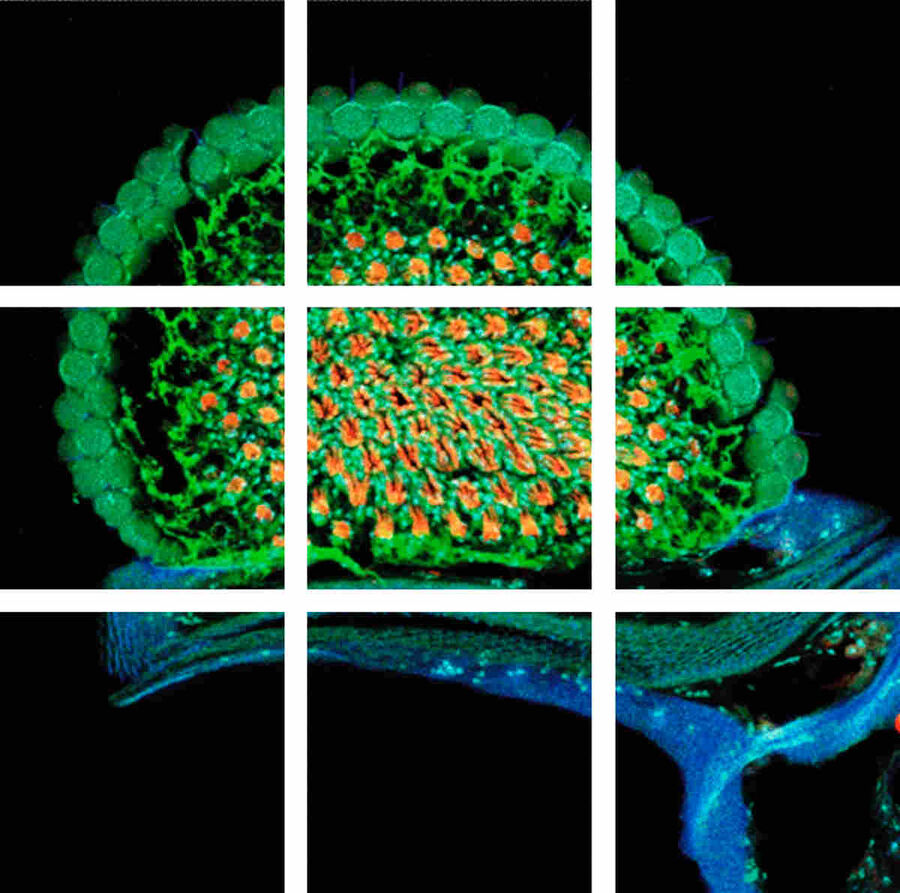
To solve this problem – large specimen, but small scanning area at high resolution – the mosaic function can be used, assembling single images to form one large super image of the specimen. The prerequisite for such a stitched super image is a tile scan, which records a defined number of adjoining single images of the sample (the "tiles"). The sample is generally moved using motorized stages with high precision. To perfectly match the tiles a small percentage (2–10 %) of overlapping pixels between single images is necessary. Most application software provides helpful functions where only two reference points at opposite corners of the specimen must be defined or the area of the specimen is automatically detected; in turn the number and position of the scan fields to cover the specimen are calculated, including predefined overlapping regions. Following the scanning process, the mosaic image is assembled using specifically designed stitching algorithms.
To solve this problem – large specimen, but small scanning area at high resolution - the mosaic function can be used, assembling single images to form one large super image of the specimen. The prerequisite for such a stitched super image is a tile scan, which records a defined number of adjoining single images of the sample (the “tiles”). The sample is generally moved using motorized stages with high precision. To perfectly match the tiles a small percentage (2-10%) of overlapping pixels between single images is necessary. Most application software provides helpful functions where only two reference points at opposite corners of the specimen must be defined or the area of the specimen is automatically detected; in turn the number and position of the scan fields to cover the specimen are calculated, including predefined overlapping regions. Following the scanning process, the mosaic image is assembled using specifically designed stitching algorithms.
3D mosaic images
The mosaic function is not only restricted to xy images, but can also be used to create large 3D images. Hence the tile scan must be performed not only in one focal plane, but in different z-planes, creating a 3D mosaic of adjacent image stacks. However, a fundamental prerequisite is a synchronized z-distance between the focal planes.
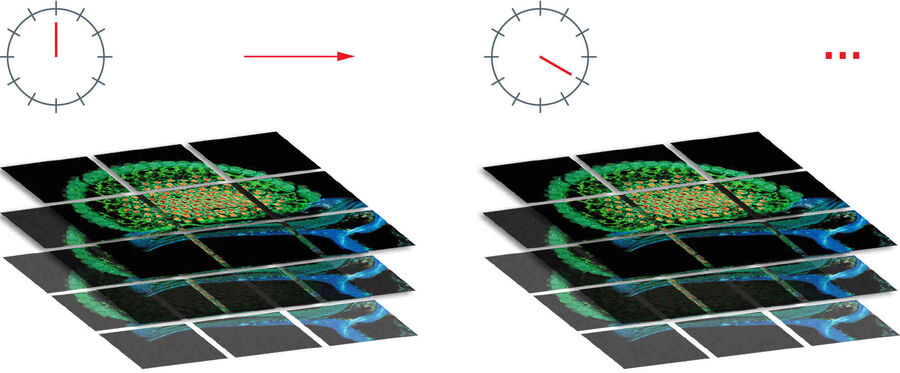
Time lapse movies
As mosaic scans can be repeated in loops, this function can be used to create time lapse movies of living samples. The only restriction would be the velocity of changes in the specimen, as tile scanning takes time, particularly in three dimensions.
Multi position function
In addition, the mosaic function can be carried out at different positions of the stage. This allows the recording of several specimens in one experimental setup (e.g. different slices on a slide or several samples in a multi-well plate). As the recording conditions will vary from specimen to specimen, the system used should provide the possibility to define different scanning setups for each position or even for each tile of the scan.
Related Articles
-
Extended Live-cell Imaging at Nanoscale Resolution
Extended live-cell imaging with TauSTED Xtend. Combined spatial and lifetime information allow…
Mar 05, 2024Read article -
How to Prepare Samples for Stimulated Raman Scattering (SRS) imaging
Find here guidelines for how to prepare samples for stimulated Raman scattering (SRS), acquire…
Feb 05, 2024Read article -
Coherent Raman Scattering Microscopy Publication List
CRS (Coherent Raman Scattering) microscopy is an umbrella term for label-free methods that image…
Sep 11, 2023Read article