FLIM imaging with STELLARIS confocal platform
This image gallery shows applications of TauSense and fluorescence lifetime imaging (FLIM) on the STELLARIS confocal platform.
3D confocal image of C. elegans neurons labelled with four red fluorescent proteins.
Image acquired with STELLARIS 8 FALCON with a single excitation line and a single detector. The 4 distinct signals were obtained with phasor separation. The use of a single laser line and single detector improves live imaging speed and lowers the laser dose. Grey: Intensity only. Colors: Merged phasor-separated signals. Scale bar: 20 μm. Tracking 4 neuronal types with 4 red fluorescent proteins on a single detector with FLIM.
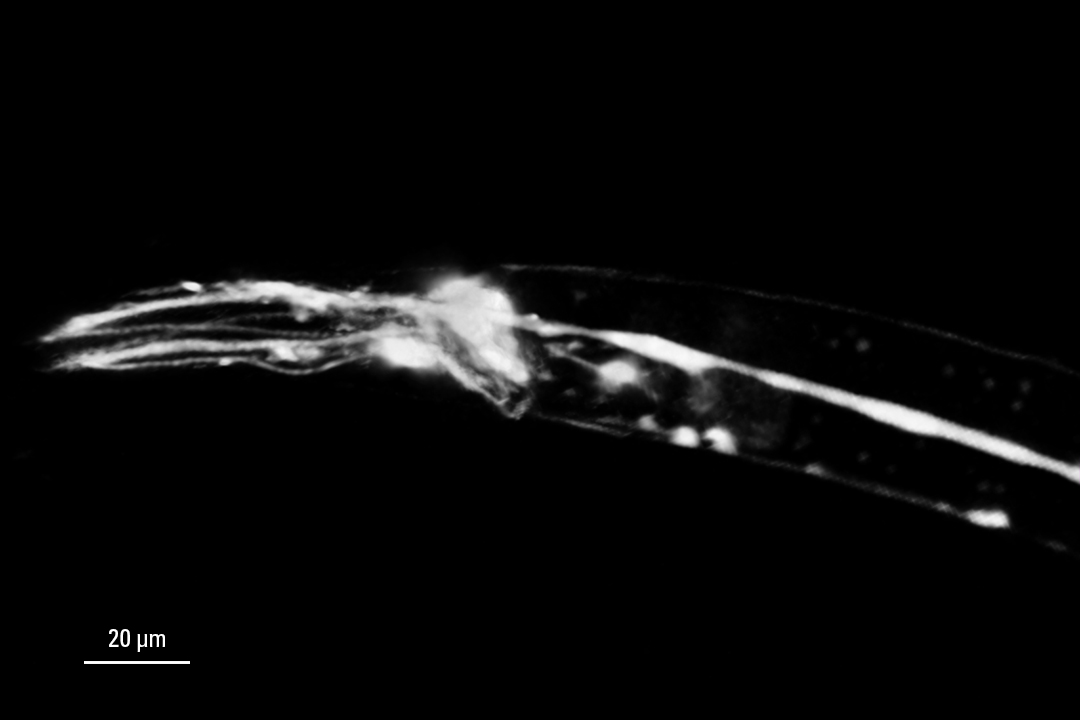
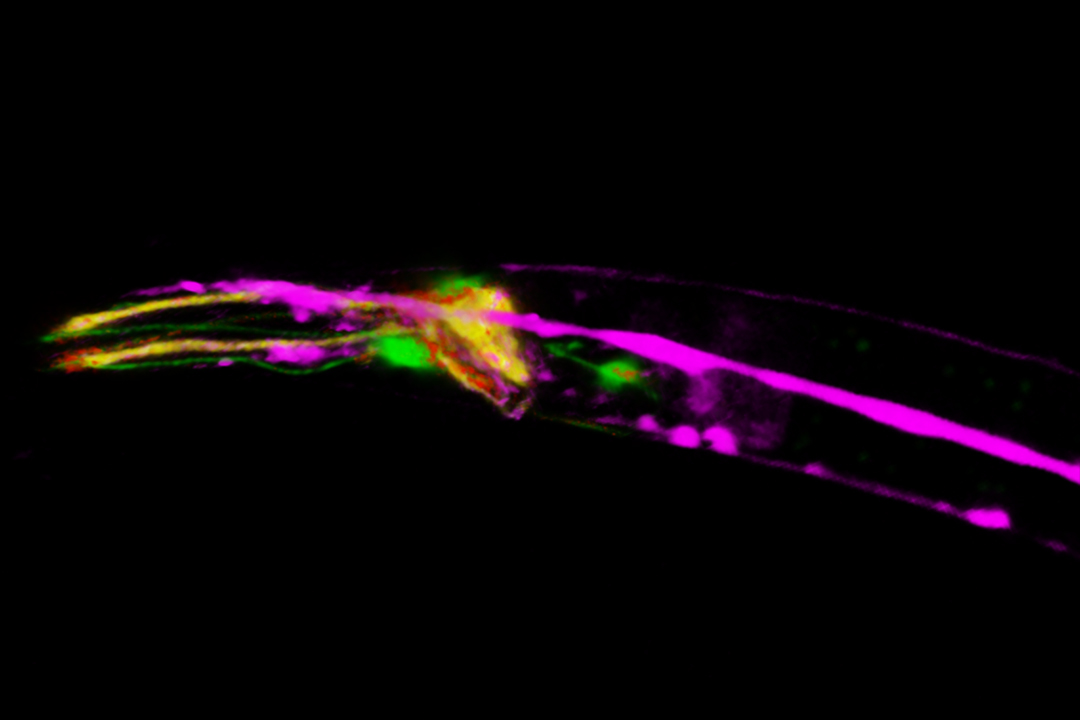
3D confocal image of C. elegans neurons labelled with four red fluorescent proteins
Transcription activity in mice gastric stem cell derived organoid
Endocytic pathway and mitochondria dynamics
Live NE-115 cells expressing mNeonGreen-LifeAct - STELLARIS
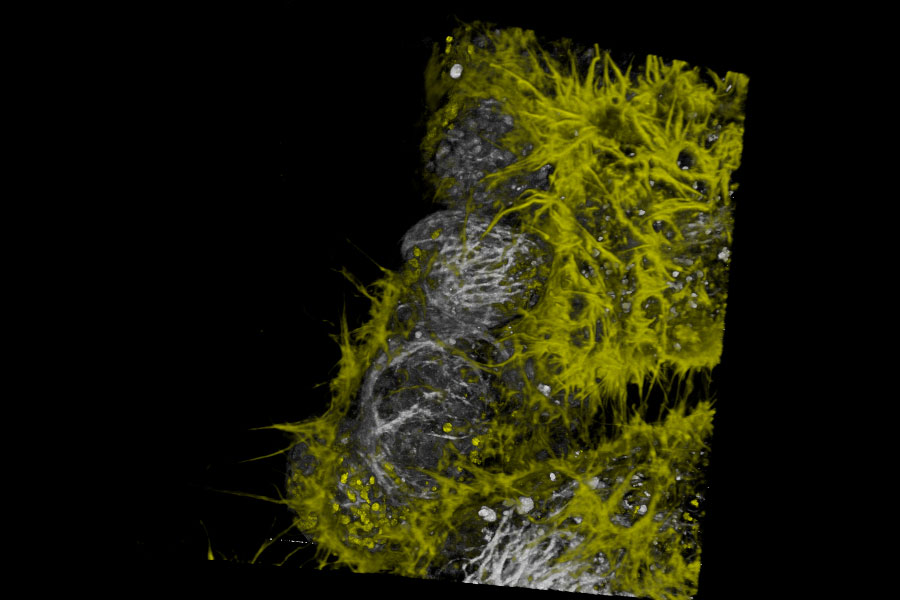
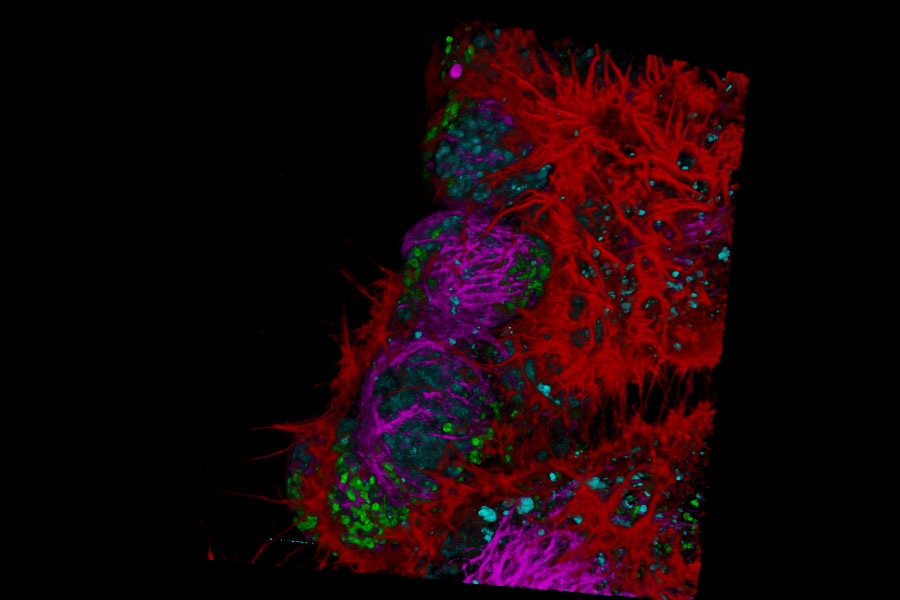
Live NE-115 cells expressing mNeonGreen-LifeAct are stained with MitoTracker Green, NucRed and SiR Tubulin. Signals are acquired with 2 detectors for two fluorophores each. Intensity image shows in yellow and gray. Signal in each detector will be distingu
Exploring Chloroplast activity with multiphoton and fluorescence lifetime imaging
Image acquired with a STELLARIS 8 DIVE FALCON.
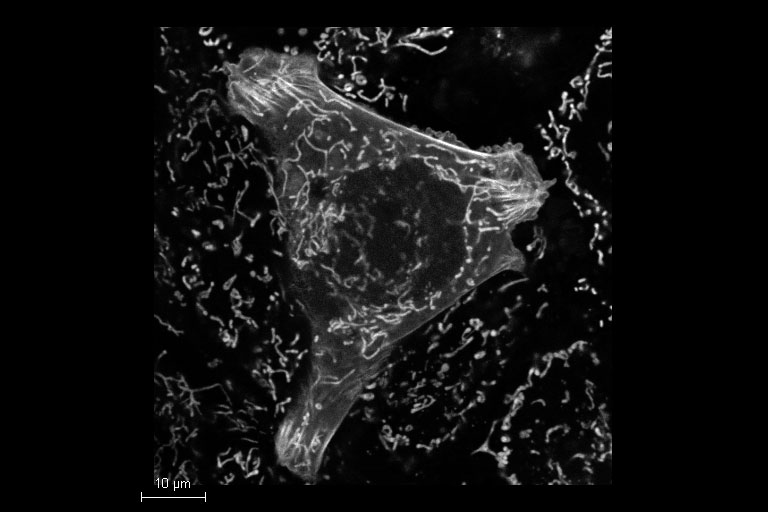
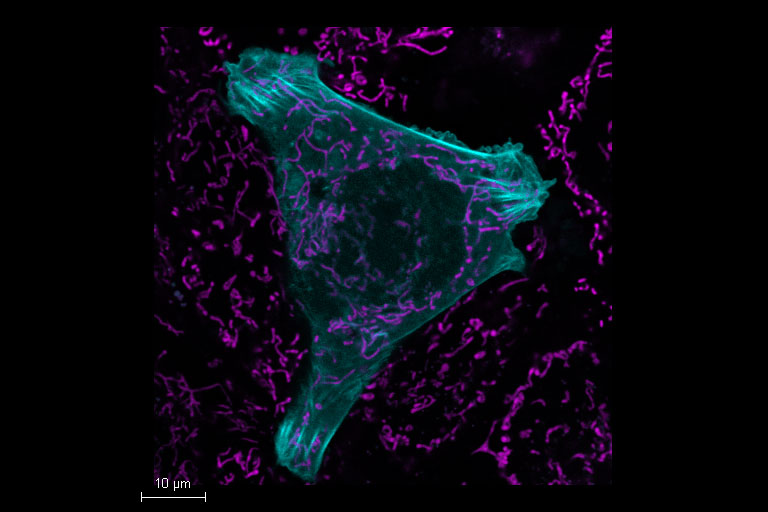
Mammalian cells expressing LifeAct-GFP (manufactured by ibidi GmbH) and labeled with a MitoTracker Green. Acquisition with one detector, separation performed by TauSeparation.
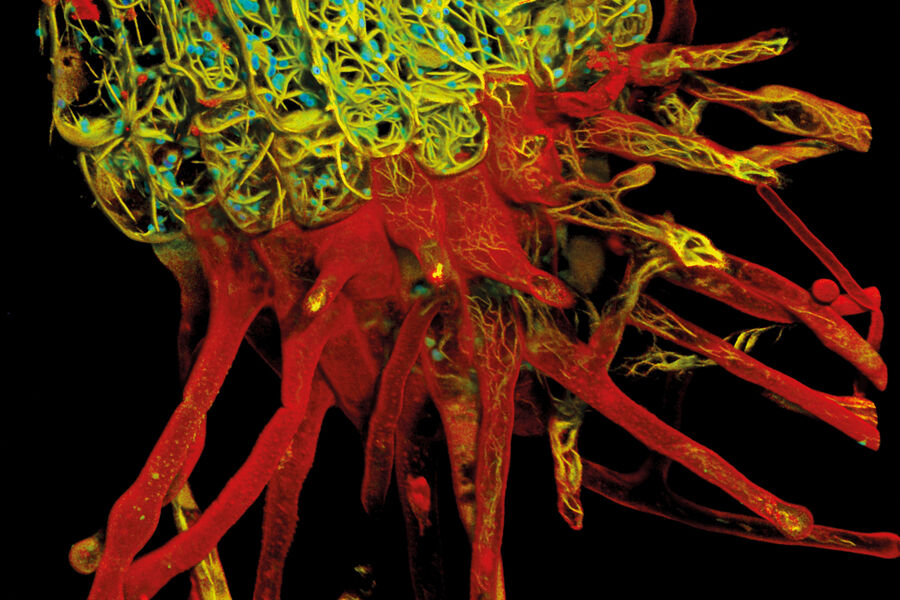
Root-hypocotyl junction of Arabidopsis thaliana
Nature is multi-dimensional. To truly comprehend its complexity, we need to look in multiple dimensions. This detailed multicolor image of Arabadopsis thaliana root junction was obtained with just one click and one detector. With conventional confocal microscopy, it would be black and white. With STELLARIS what we see is transformed by an extra dimension of lifetime information. Not only is the remarkable architectural diversity of the actin-containing root system revealed with more clarity, but also its relationship to the cell wall and chloroplasts—structures that would be difficult or impossible to distinguish using the intensity data alone. The result is not just a snapshot of nature’s beauty, but an information-rich image that progresses our understanding of plant biology.