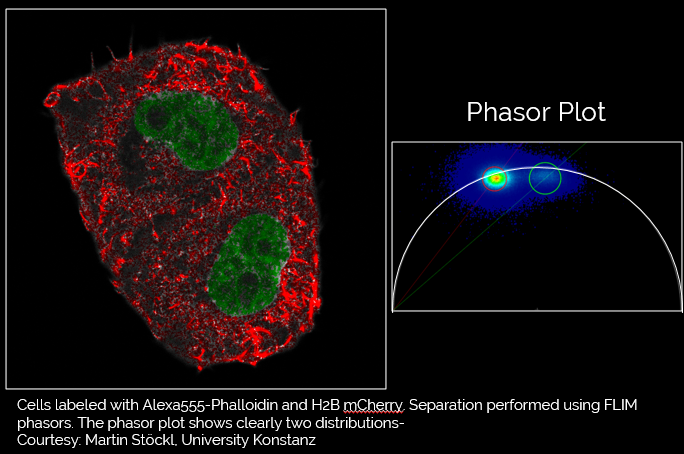
How is the phasor plot generated?
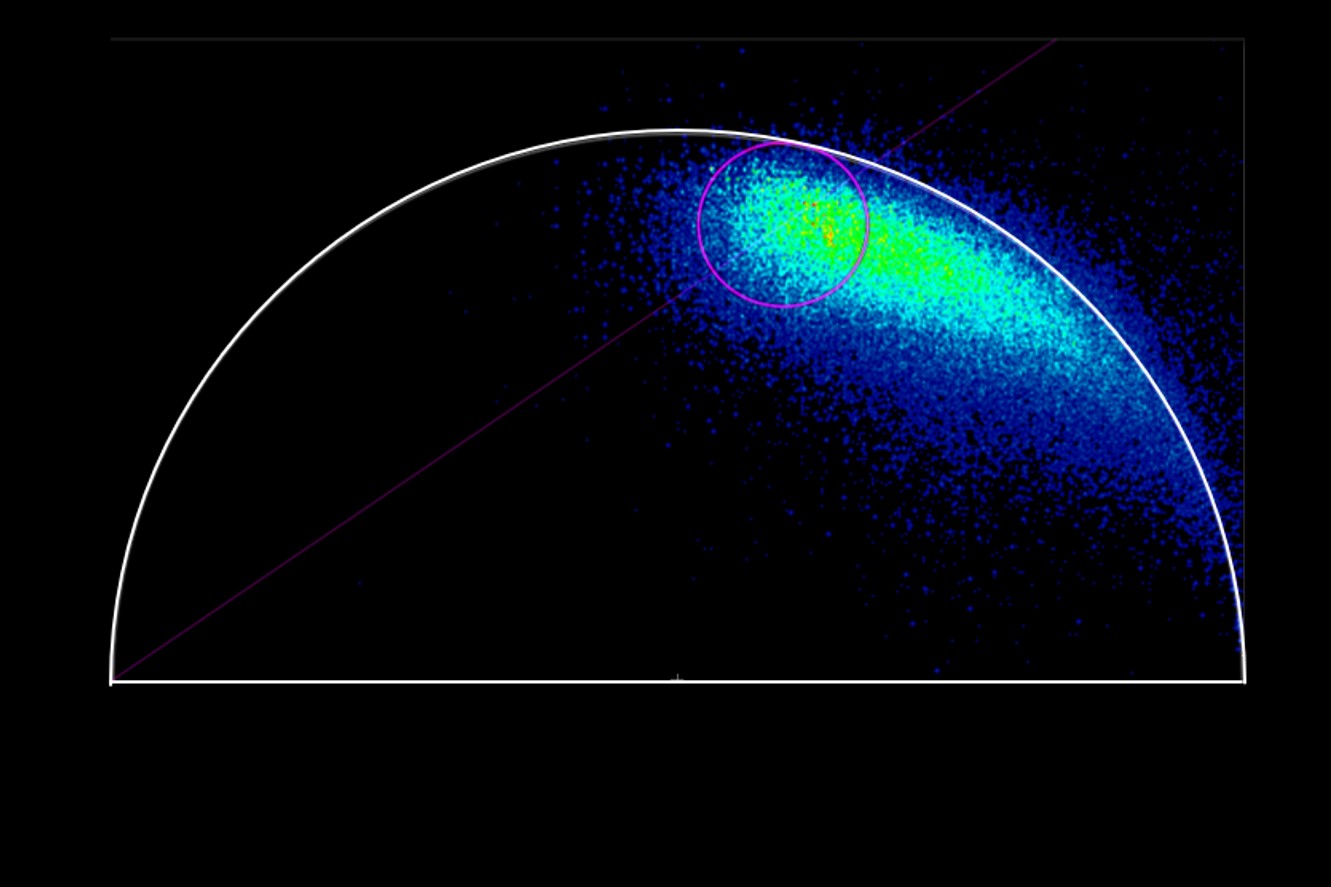
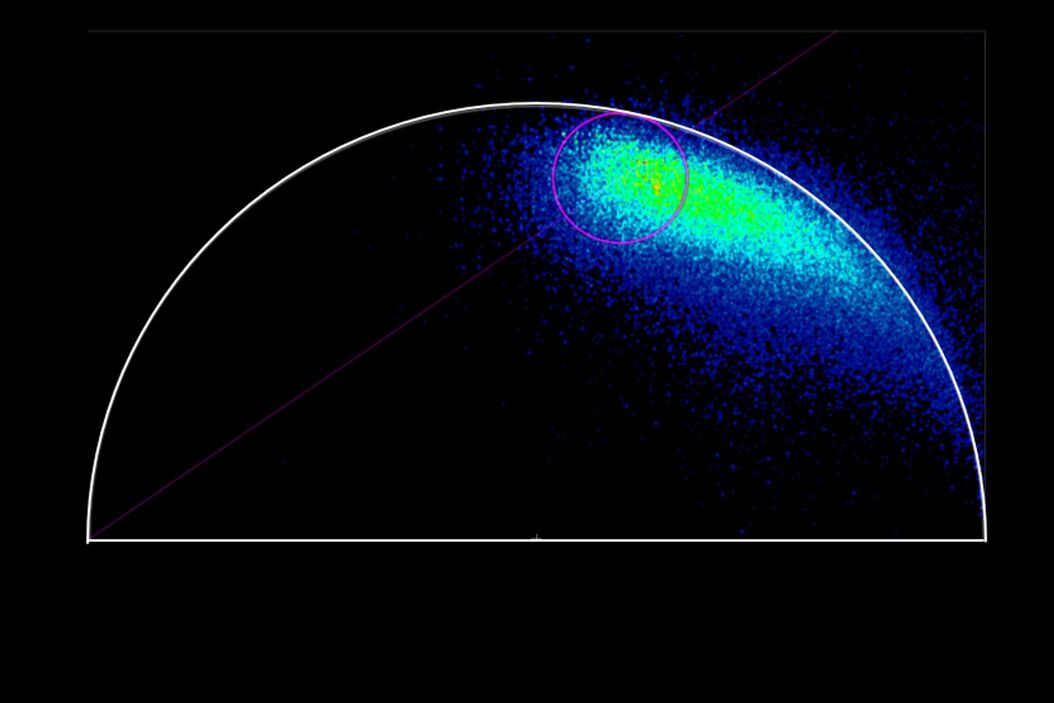
When the data is acquired with Time Resolved Single Photon Counting (TCSPC) systems, the phasor FLIM distributions are derived from a Fourier transform (Digman et al., 2008). Each pixel in the image corresponds to a point in the phasor plot.
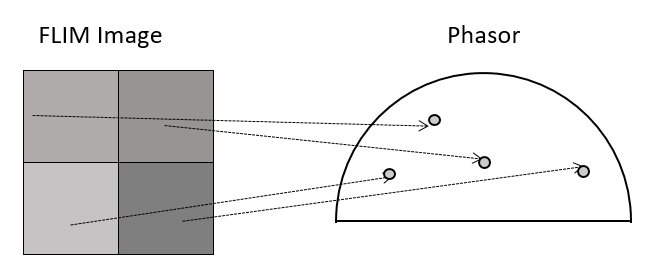
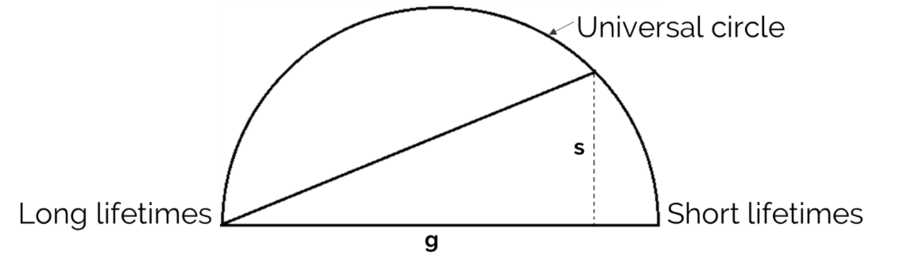
Rules: the algebra of phasors
There are only a few rules in the phasor approach and they enable a straightforward interpretation of the lifetime distribution. Three of the most important ones are shown below (E. Gratton, 2018).
1. Single exponential lifetimes lie on the universal circle line.
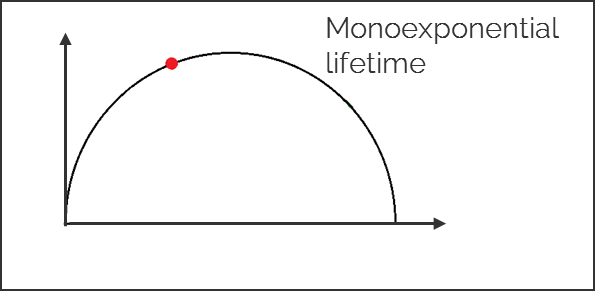
2. Multi-exponential lifetimes are found inside the universal circle and are a linear combination of their single exponential lifetime components.
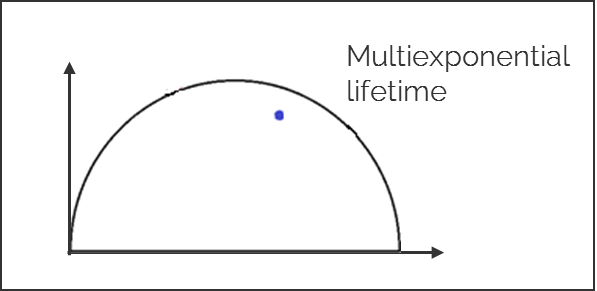
3. The ratio of the linear combination for a multi-exponential lifetime (experimentally measured) determines the fraction of the components (f1 and f2) from their two single exponential lifetimes (τ1 and τ2).
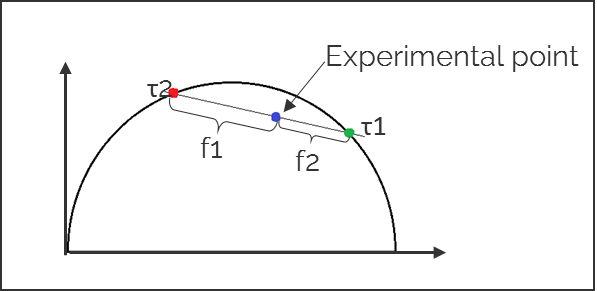
Advantages of using phasors
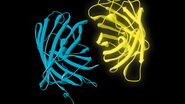
Phasor FLIM is a very powerful analysis tool for molecular species separation and FRET analysis, in particular when the donor has a multi-exponential lifetime, something which is typical of CFP [cyan fluorescent protein] (Caiolfa et al., 2007). Moreover, phasor FLIM combined with STED, allows you to use less STED power to reach the same resolution (Lanzanò et al., 2015).
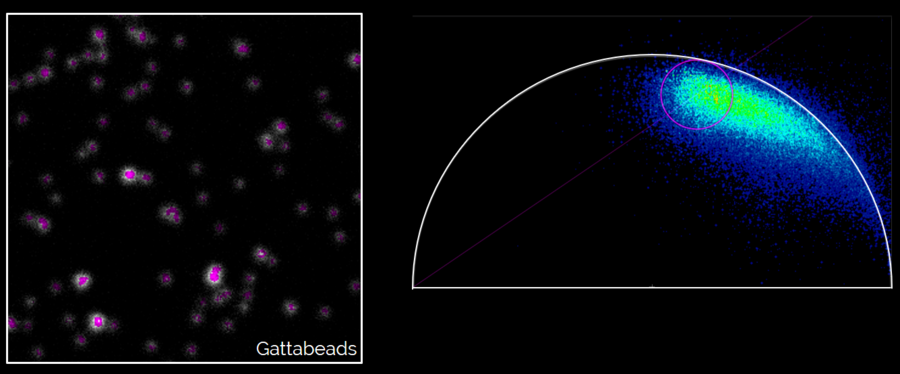
Left: Gattabeads imaged with SP8 STED system having FLIM capability. Right: By highlighting the long lifetime pixels in the phasor plot, the resolution of the STED image is improved.