Stalled by the limitations of probes
A drawback of many methods for metabolic imaging is their reliance on labels to track metabolic changes. Macro-level methods, like PET, use radioactive labels or tracers. Microscopy methods often rely on fluorescent probes. Unfortunately, fluorescence intensity measured from probes can be inconsistent – values depend on how data are collected, on the concentration of the probe and on how well the probe penetrates a sample, which may require fixation and perfusion. Furthermore, some probes are toxic to cells, may bind non-specifically, or can destroy biochemical and physiological conditions in tissues. Such traits impact results and make repeat measures impossible. Fluorescence lifetime imaging, or FLIM, offers an alternative: label-free metabolic imaging as a workable and minimally invasive tool to examine metabolism patterns.
A new dimension for metabolic imaging
In contrast to measuring the intensity of light emitted by a fluorophore returning to its ground state after excitation, FLIM records the average time that the fluorophore remains in the excited state. That time interval is a unique property of the fluorophore that is influenced by configuration changes to the fluorophore through binding or interactions with other molecules.
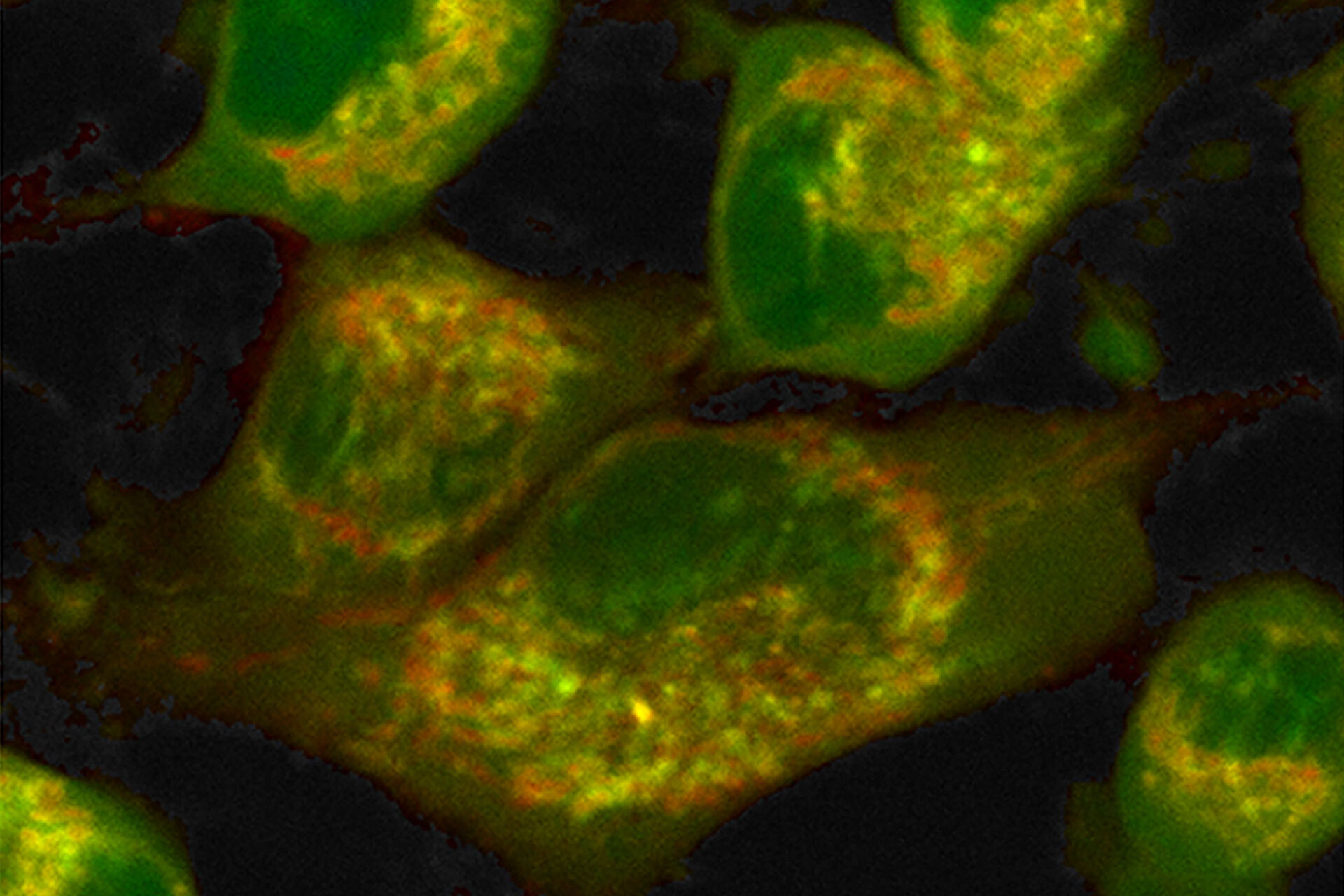
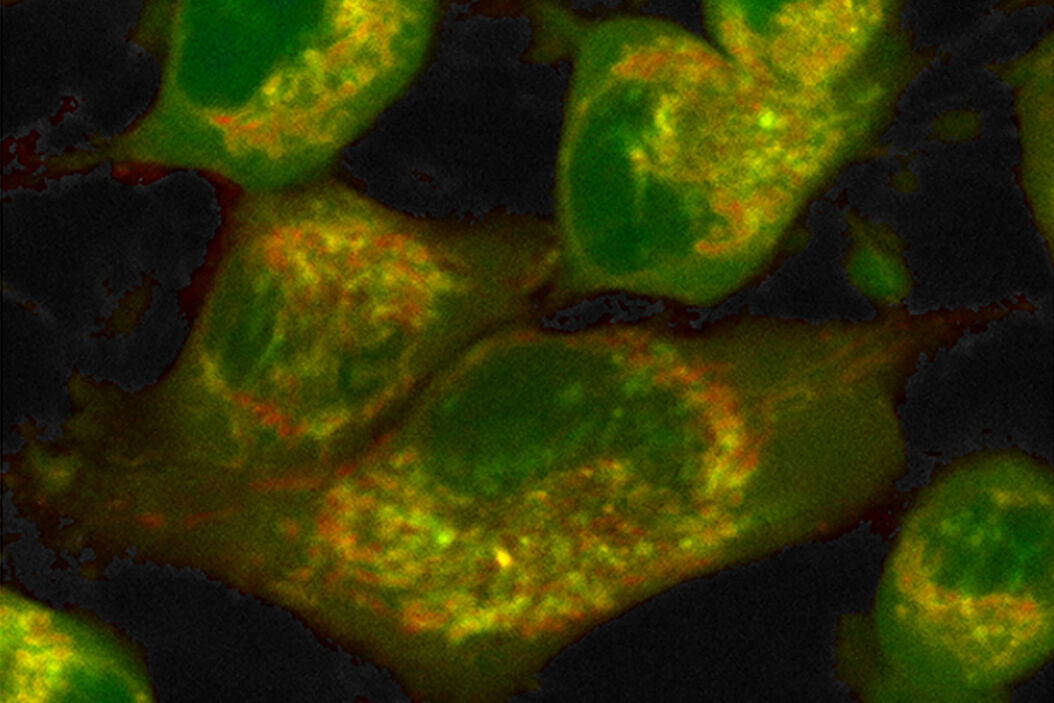
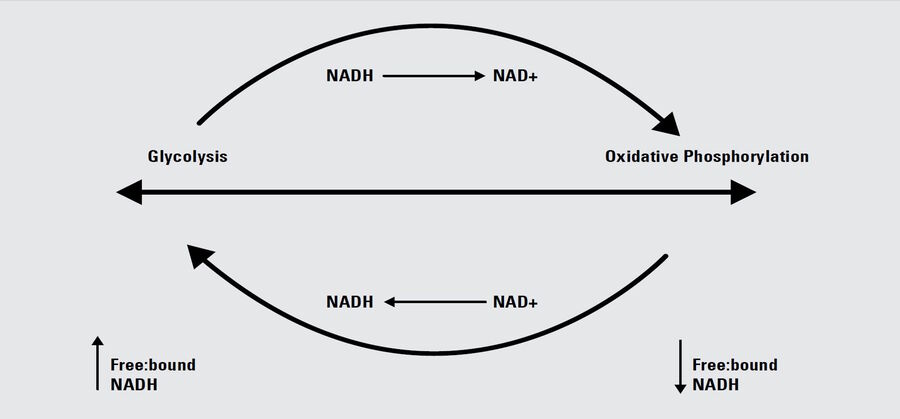
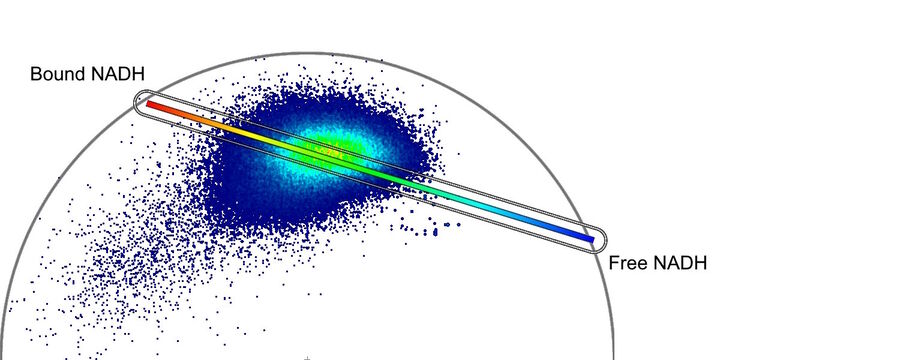
Label-free metabolic imaging via FLIM uses autofluorescent metabolic enzyme cofactors, like NADH. Cellular NADH levels follow changes in metabolism. A cell favoring glycolysis has a higher ratio of NADH to its oxidized counterpart, NAD+, than a cell preferentially performing oxidative phosphorylation. That reduction-oxidation ratio, conversely, correlates with the ratio of free to protein-bound NADH, a configurational difference that FLIM can detect. The fluorescence lifetime of free NADH (~0.4 ns) is significantly lower than that of protein-bound NADH, providing functional information about the metabolic status of a cell along a phenotypic spectrum from glycolysis at one end, where the free:bound NADH ratio is high, to oxidative phosphorylation at the other end, where the free:bound NADH ratio is low (Figure 1).
Combined with a multiphoton microscope like STELLARIS 8 DIVE, FLIM has advantages for label-free metabolic imaging of live cells and thick tissue sections. A multiphoton microscope excites fluorophores with infrared wavelengths rather than the ultraviolet light of single-photon excitation that can cause phototoxicity in live cells. The great focal depth but small focal volume of a multiphoton microscope facilitates deep tissue imaging with minimal out-of-focus fluorescence, complementing the sensitivity of FLIM in recording the low-level signals of autofluorescent species at high resolution and precision. Finally, fluorescence lifetime is unaffected by probe concentration, photobleaching, light scattering or excitation light intensity – it is an orthogonal dimension liberated from constraints of intensity measures that expands information extracted from images.
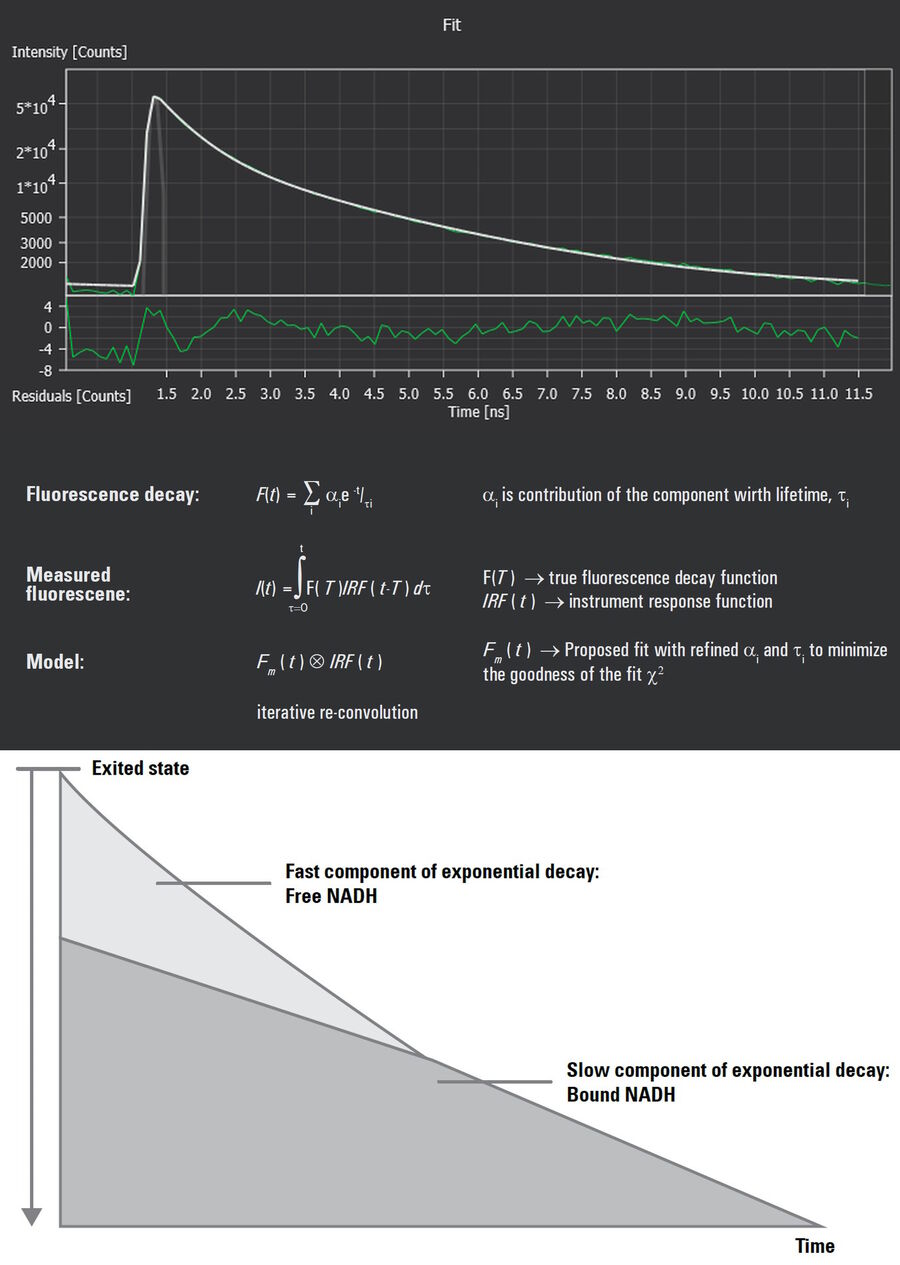
TauContrast, a TauSense imaging tool available on STELLARIS 8 DIVE, generates images from the average photon arrival time and the number of photon counts from each pixel (intensity). The outcome is a detailed visualization of changes in tissue microenvironment that reflect metabolic state. However, TauContrast does not quantify fluorescence lifetime and thus, cannot report a free:bound NADH ratio. Quantification is done by fitting a mathematical model to a histogram of photon arrival times that describes the exponential decay of the fluorescence lifetime of NADH (Figure 2). The curve is a composite of two exponential decays (from free and bound NADH), skewed by variation and the instrument response function Thus, this complex quantification method requires the trained eye of an expert. But there is a better way.
Fluorescence lifetime estimation by curve fitting
Untethering metabolic imaging from complex modeling
Phasor analysis grants non-experts quick and easy access to quantifiable metabolic information by eliminating the need to model complex decay exponentials. In phasor analysis, fluorescence lifetime signals from every pixel in an image are Fourier transformed into frequency components – a phasor – that are charted on a phasor plot (Figure 3). The power of phasor analysis lies in its interpretability based on negative and positive controls. Those controls fall on the outer edge, or “universal circle,” of a phasor plot. In the case of NADH-based metabolic imaging, the controls are pure free NADH and pure protein-bound NADH. The phasor of each pixel in an image emerges from a mixture of those two states of NADH and thus, falls on a line between the two controls. That line is called the “metabolic trajectory.” The concentration of free NADH relative to bound NADH is thus calculated for every pixel by looking at the position of that pixel’s phasor on the plot. Going one step further, the average phasor of all pixels that make up a cell in the image is the cell’s phasor fingerprint and it characterizes the state of that cell along that metabolic trajectory.
With phasor analysis, no mathematical modelling is necessary, and results are insensitive to probes, making metabolic imaging accessible to any lab, regardless of expertise. The phasor plot is also a robust and intuitive visualization of the phasor fingerprint of a chemical or of a cell, which makes presenting and comparing results easier. Finally, a selection of interest on a phasor plot can be mapped back on the image to localize those signals to cells or subcellular regions.
As a fully integrated multiphoton microscope system for FLIM, STELLARIS 8 DIVE FALCON augments the power of FLIM with advanced microscopy features that allow tile-scanning and 3D acquisition at video-rate frame speeds. Crucially, STELLARIS 8 DIVE FALCON transforms FLIM applications like metabolic imaging from complex specialty techniques to widely available, easy-to-use tools for collecting and interpreting FLIM data via phasor analysis. The following studies exemplify the use of phasor analysis to link metabolism and cell behavior.
Metabolic status predicts differentiation potential and direction
In a series of articles, Stringari and colleagues use FLIM and phasor analysis to characterize the metabolic and differentiation state of germ cells in Caenorhabditis elegans [1], of neural progenitor and stem cells (NPSCs) [2], and of epithelial cells in small intestine crypts [3]. In all three studies, the phasor fingerprint of individual cells correlate with the differentiation state of the cell and its commitment to a particular differentiation path. In C. elegans, distinct clusters of cell phasor fingerprints align with a gradient of germ cells that transition from an undifferentiated “stem cell-like state” distally to more differentiated cells proximally [1]. Undifferentiated NPSCs exhibit a glycolytic phenotype with high free:bound NADH ratios, whereas differentiated neurons lean toward oxidative phosphorylation characterized by low free:bound NADH ratios. Furthermore, NPSCs can be classified as neural progenitors or glial progenitors based on phasor fingerprint, as the latter reside further on the oxidative phosphorylation side of the metabolic trajectory [2]. A similar alignment of metabolic and differentiation state is apparent in epithelial cells lining the crypts of small intestine tissue. Stem cells at the bottom of the crypt are distinctly glycolytic. The increasingly differentiated cells lining the crypt to the surface have phasor fingerprints that progressively shift toward oxidative phosphorylation [3].
Metabolic imaging as a tool for high-throughput drug screening
A system to non-destructively track the metabolic response of live cells to external stimuli may constitute an effective tool for in vitro high-throughput drug screening. In a 2016 study, Datta and colleagues provide proof-of-concept to use FLIM and phasor analysis for that purpose [4]. The team triggered hypoxia in cardiomyocytes by restricting culture oxygen levels or exposing cells to an anticancer drug (cisplatin) or an antiviral (AZT) known to produce oxidative stress. All three treatments resulted in higher free:bound NADH ratios in phasor plots corresponding to a metabolic shift towards glycolysis similar to subjecting cells to KCN, a poison that shuts down oxidative phosphorylation.
Understanding methionine metabolism in cancer cells
A majority of malignant cancer cells exhibit a methionine-dependent metabolic phenotype. Unlike non-tumorigenic cells, they cannot proliferate in growth media where methionine is replaced with homocysteine.
Borrego and colleagues used FLIM and phasor analysis to examine differences in metabolism between a breast cancer cell line and a derived clone line shown to be resistant to methionine dependency and to have lost the transformation phenotype typical of tumorgenicity [4]. Grown on methionine medium, both cell lines exhibited similar metabolic profiles with approximately 80% of the NADH signal representing a low free:bound NADH ratio. Transferred to homocysteine medium, the resistant clone line shifted further toward glycolysis, with only 64% low free:bound NADH ratio compared to 70% in the original cells. The resistant clones may be less reliant on oxidative phosphorylation and thus more tolerant of methionine stress.
Detecting biomarkers of oxidative stress
FLIM with phasor analysis also detects oxidized lipids which appear as long lifetime species (LLS) on a phasor plot [6]. These LLS are biomarkers of oxidative stress and appeared in the cardiomyocytes studied by Datta et al. upon exposure to hypoxia and stress-inducing drugs [4]. In a similar fashion, the methionine-dependent cancer cell lines in the study by Borrego et al. showed heightened generation of LLS [5]. Under normal growing conditions, the clone line resistant to methionine dependency had a higher average fraction of LLS compared to the original cell line (22 vs. 7%). However, upon transfer to homocysteine medium, that fraction increased by 29% in the original cell line and only 5% in the clone line, hinting at a possible protective function of LLS. Both studies put forward an oxidative stress trajectory visible on the phasor plot as a method to examine the role of oxidation in cancer, atherosclerosis, and heart disease.
Summary and conclusion
Label-free FLIM with phasor analysis has opened intriguing possibilities to explore the dynamic environment and function of cells and tissues. Free of probe limitations and complex model fitting, FLIM with phasor analysis also goes far beyond NADH. Stringari documented unique phasor fingerprints of other intrinsic fluorophores, including GFP, FAD, retinol, retinoic acid, collagen and protoporphyrin IX [1]. By streamlining FLIM data collection and interpretation via phasor analysis, STELLARIS 8 DIVE FALCON easily transforms those phasor fingerprints into quantifiable markers to track changes in metabolism, 3D microenvironment and signaling pathways in single cells or across tissues. Furthermore, Cao and colleagues demonstrate the simultaneous measurement of two intrinsic fluorophores despite spectral bleed-through which greatly expands analytical possibilities [7]. FLIM with phasor analysis is a powerful and robust method to examine biochemical and physiological dynamics at the cellular and subcellular level. A methodology previously limited to expert microscopists with experience in complex experimental setups and data modelling, STELLARIS 8 DIVE FALCON now makes metabolic imaging a research tool for every scientist, every lab and every idea.