Live cell imaging with STELLARIS confocal platform
The STELLARIS platform provides you with solutions whether you need long acquisition times for high-resolution 3D reconstructions or the highest frame rates to capture rapid dynamic events.
Zebrafish heart muscle cells and blood flow in transmission
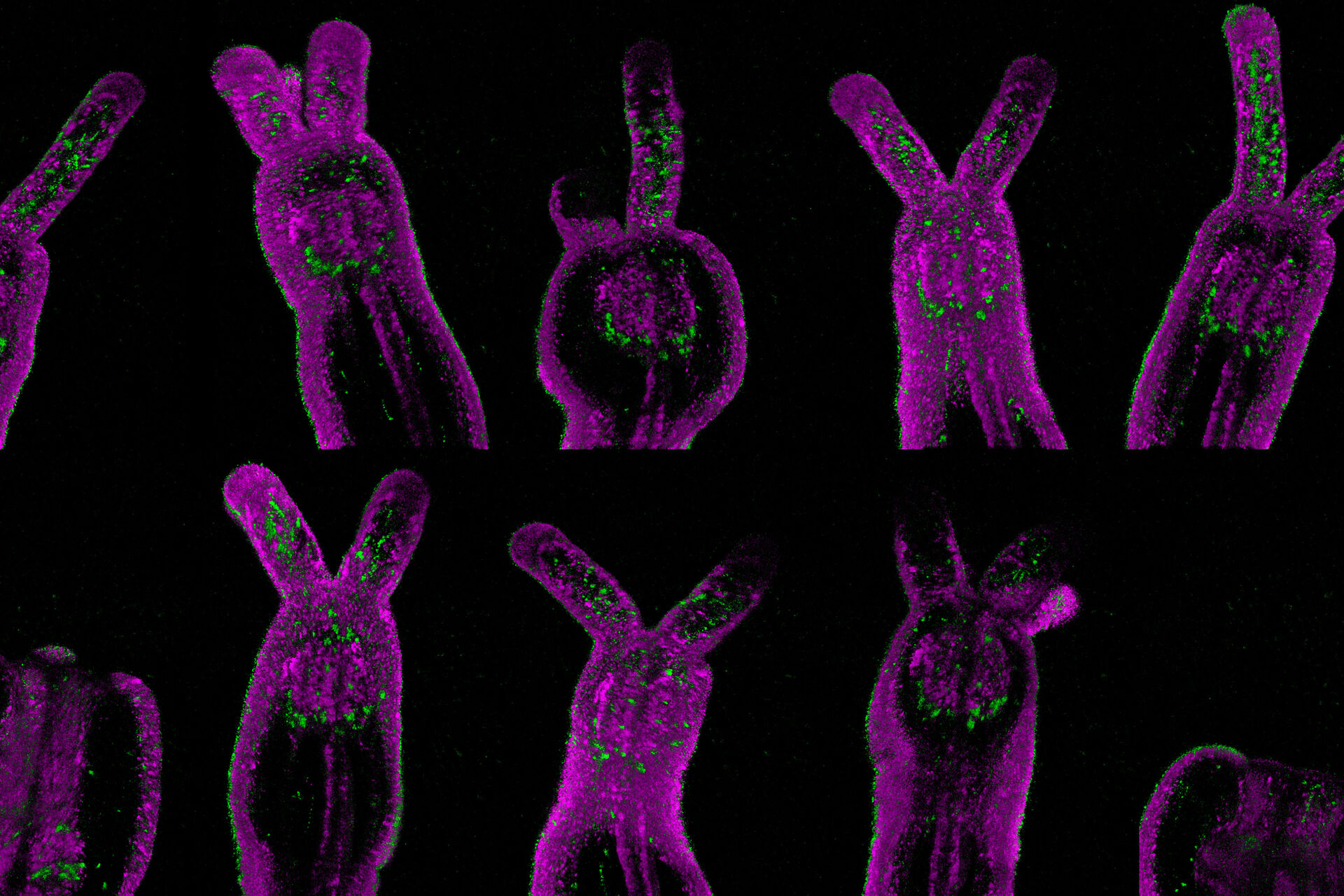
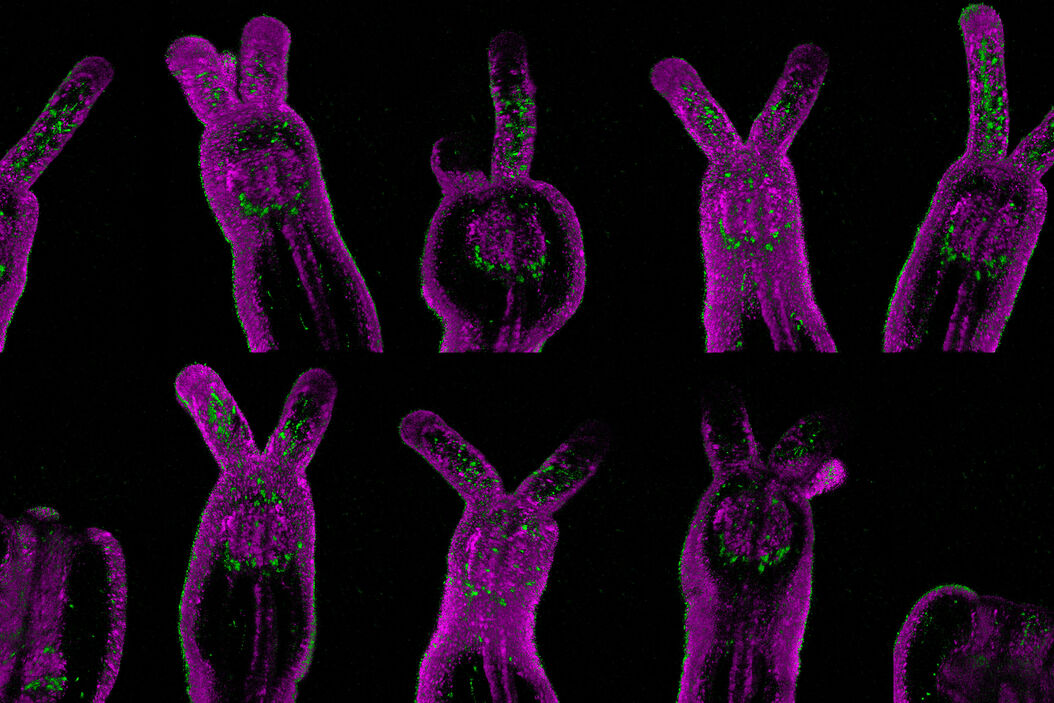
Transcription activity in mice gastric stem cell derived organoid
Gentle 3D live imaging of Hydractinia symbiolongicarpus
Endocytic pathway and mitochondria dynamics
3D live Imaging of Spirogyra sp.
Zebrafish heart muscle cells and blood flow in transmission
Volumetric image of a Zebrafish beating heart
Exploring Chloroplast activity with multiphoton and fluorescence lifetime imaging
Image acquired with a STELLARIS 8 DIVE FALCON.
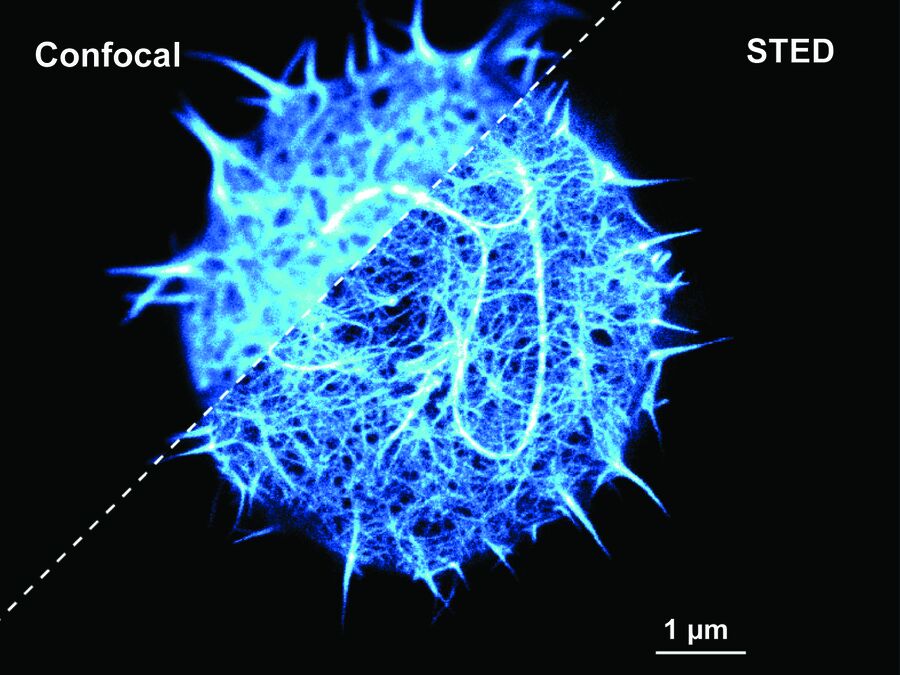
Cytoskeleton and membranes in live cell imaged with TauSTED
Image acquired with STELLARIS STED.


Cytoskeleton and membranes in live cell imaged with TauSTED.
3D confocal image of C. elegans neurons labelled with four red fluorescent proteins.
Image acquired with STELLARIS 8 FALCON with a single excitation line and a single detector. The 4 distinct signals were obtained with phasor separation. The use of a single laser line and single detector improves live imaging speed and lowers the laser dose. Grey: Intensity only. Colors: Merged phasor-separated signals. Scale bar: 20 μm. Tracking 4 neuronal types with 4 red fluorescent proteins on a single detector with FLIM.
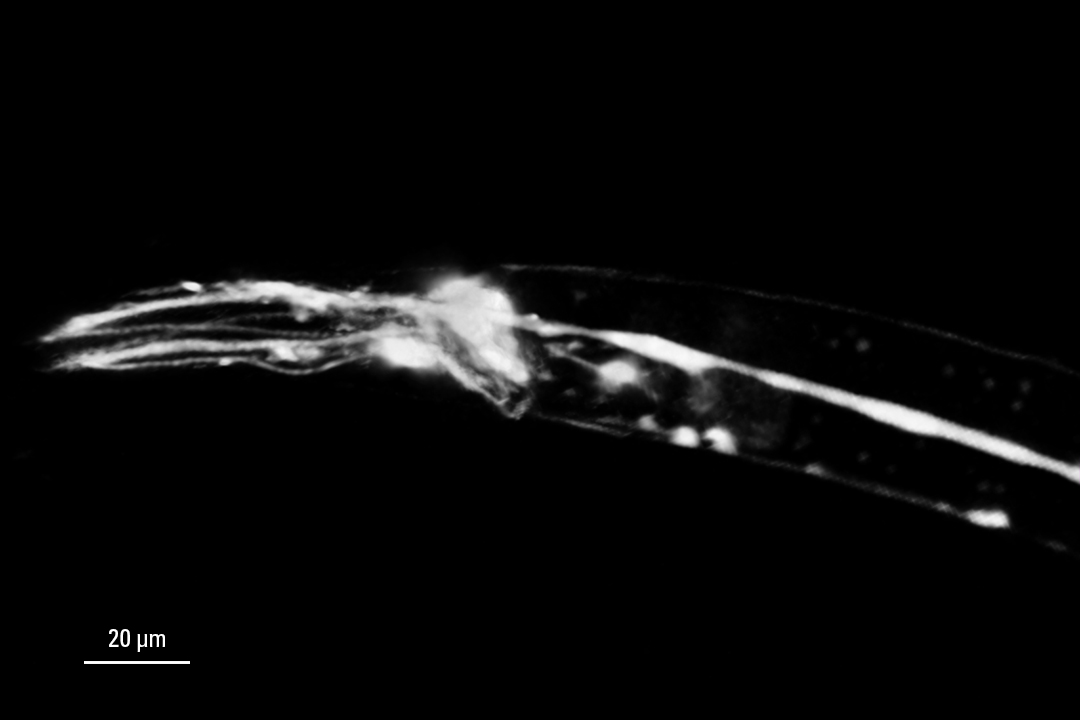
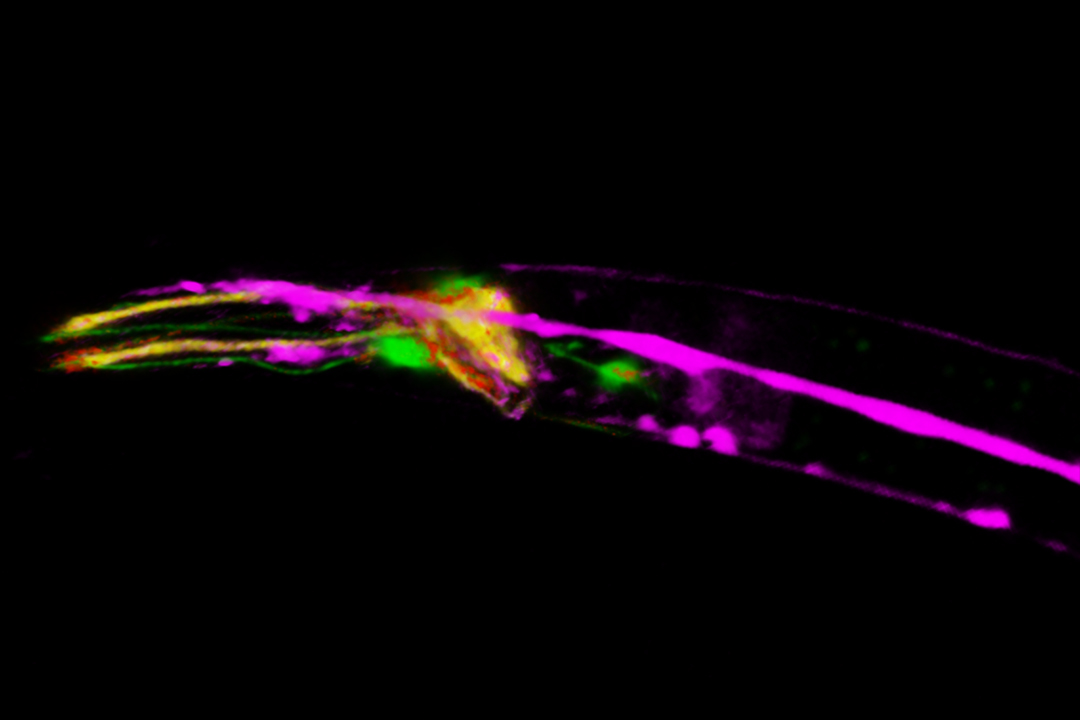
3D confocal image of C. elegans neurons labelled with four red fluorescent proteins
Cytoskeleton and membranes in live cell imaged with TauSTED
Image acquired with STELLARIS STED. Live cell TauSTED on U2OS cells, using labels for actin (SiR-actin, glow), microtubules (SPY555-tubulin, cyan), and membranes (CF488A coupled to WGA, green). SiR and SPY are available from Spirochrome. CF dyes are available from Biotium Inc. Scale bar: 10 μm
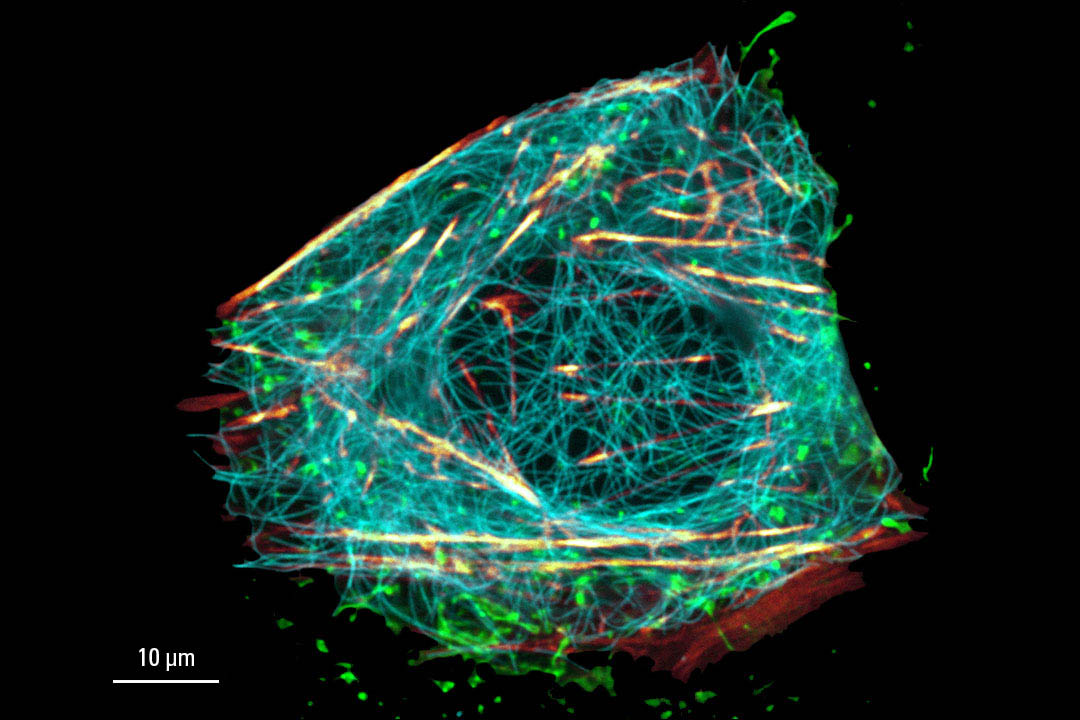
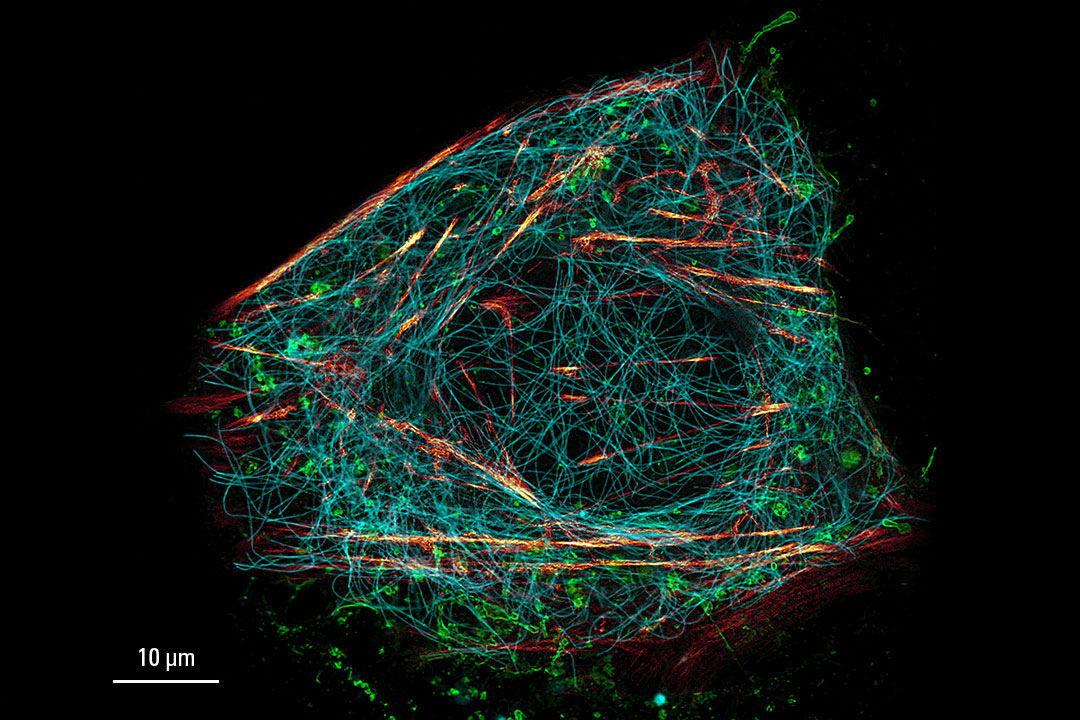
Cytoskeleton and membranes in live cell imaged with TauSTED
Calcium waves in mammalian cells
Lifetime-based multiplexing in live cells using TauSeparation
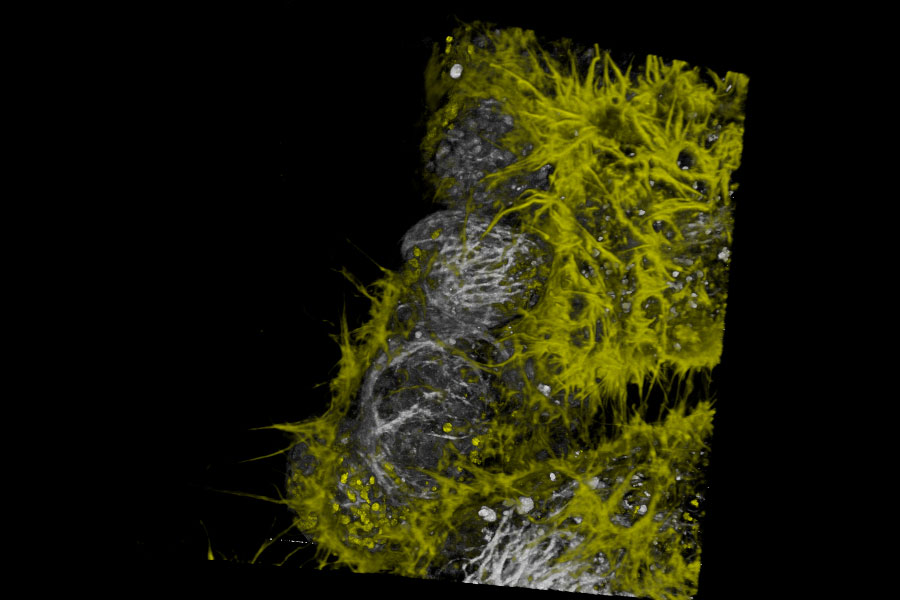
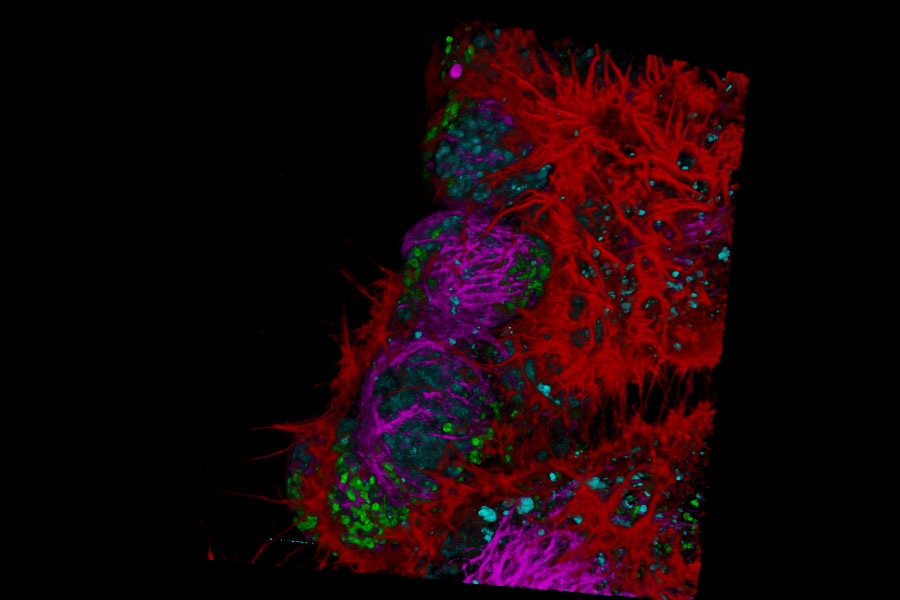
Live NE-115 cells expressing mNeonGreen-LifeAct are stained with MitoTracker Green, NucRed and SiR Tubulin. Signals are acquired with 2 detectors for two fluorophores each. Intensity image shows in yellow and gray. Signal in each detector will be distingu
Live T-cell imaged with STED