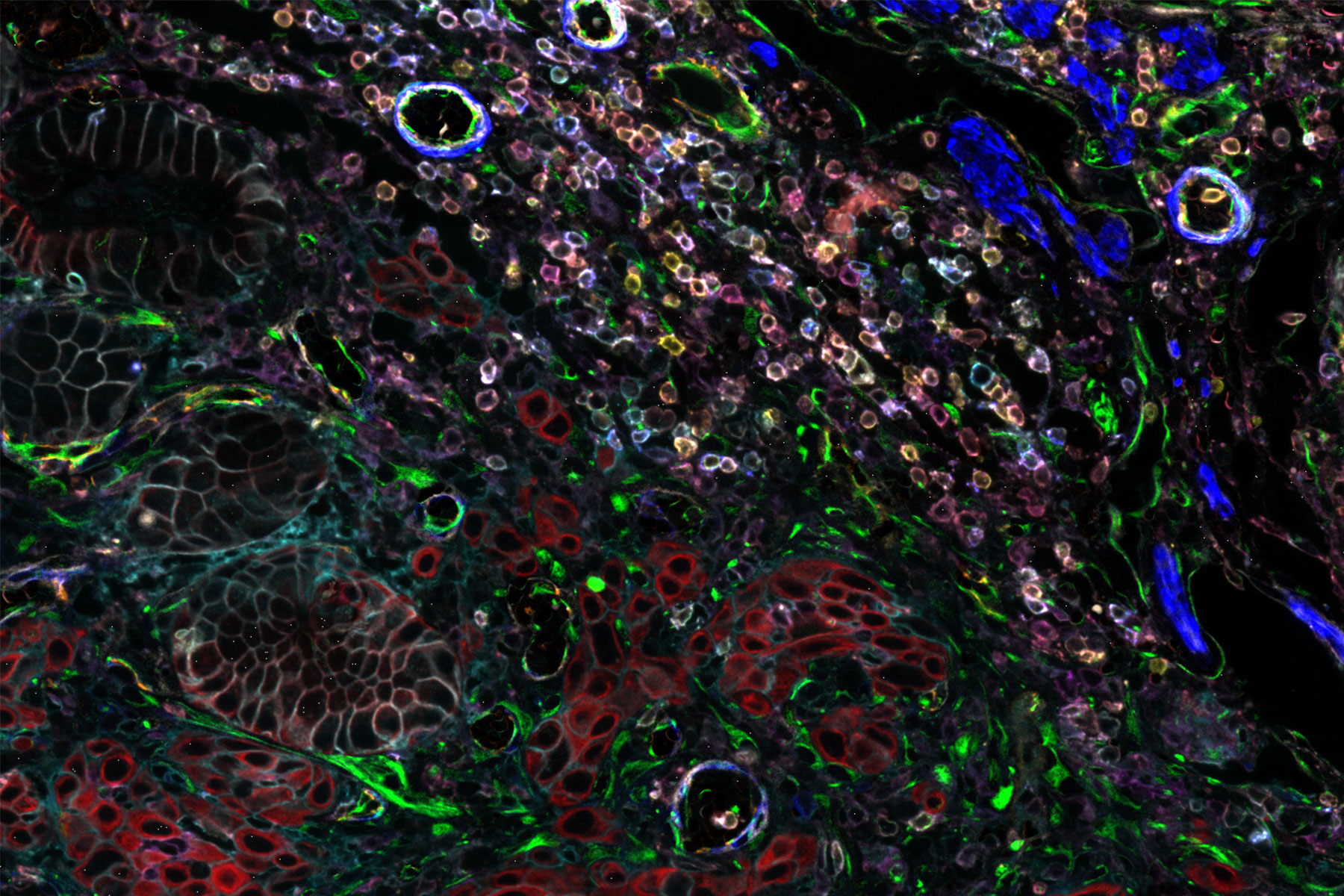
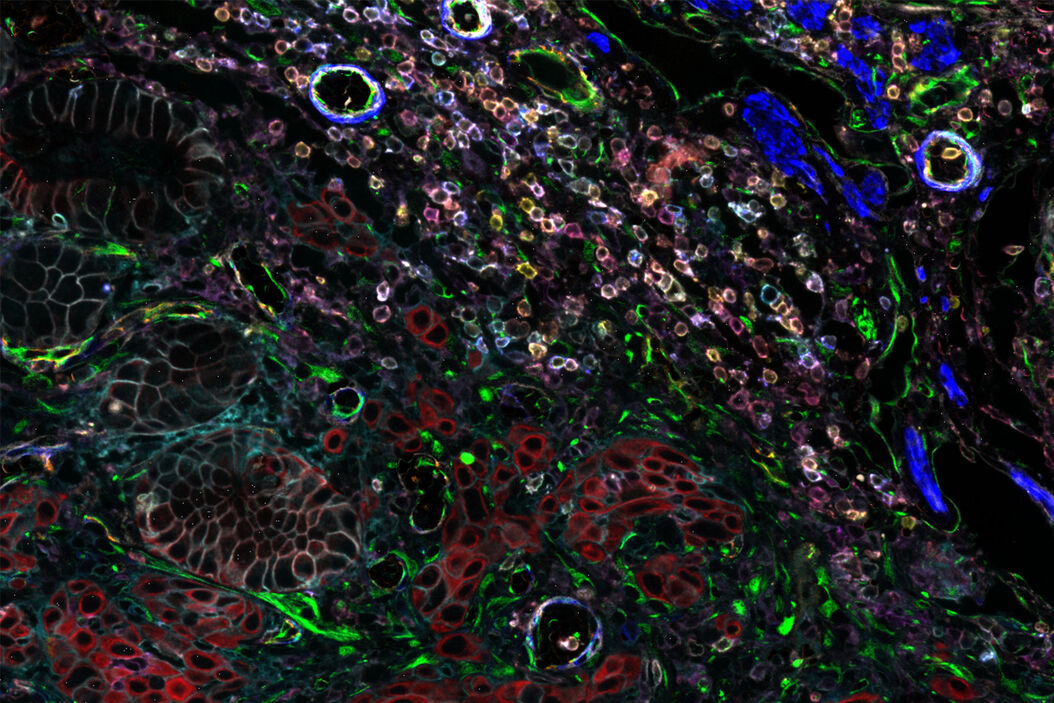
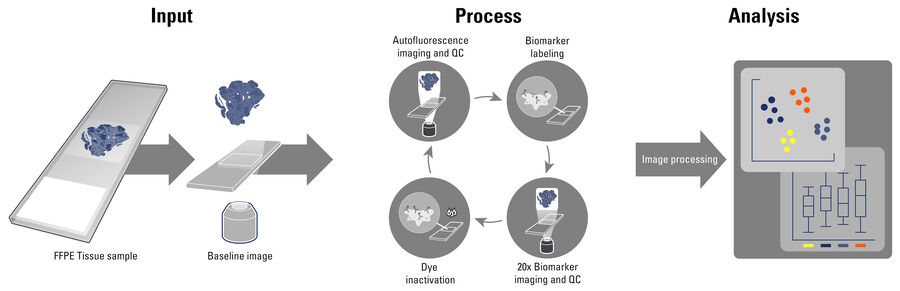
Cell DIVE is an antibody-based hyperplexed imaging technique to visualize and analyze spatial cell biology and function. Formalin-fixed, paraffin-embedded (FFPE) samples as whole slides, a region of interest, or tissue microarrays (TMA) are stained with antibodies directly labeled with cyanine-based probes, such as CyDye™ or Alexa Fluor dyes. A patented dye-inactivation process allows repeated staining and imaging of samples to create accurate, high-resolution, and comprehensive views of 60+ biomarkers in a single tissue section (Figure 1).
Optimal antibodies – cornerstone of precise multiplexed imaging analysis
Pivotal to the accuracy of any multiplexed imaging is the use of highly sensitive and specific antibodies that deliver a clear signal when and where a target protein is expressed – and only then. For Cell DIVE, those antibodies must be compatible with a two-step antigen retrieval process that appropriately unmasks a wide variety of antigens in FFPE samples, and with the gentle dye-inactivation process.
Identifying antibodies with these features involves a systematic, multi-step testing process that starts with the identification of antibody candidates in the literature and culminates in:
- The clear validation that an antibody performs robustly in the Cell DIVE workflow
- The establishment of optimal staining conditions
- The compilation of key data for sourcing, information on antigen effects due to dye inactivation, and guidance on dye loading ratio for direct conjugation or an indication to use secondary labeling
Confidence in accurate results with validated Cell DIVE antibodies
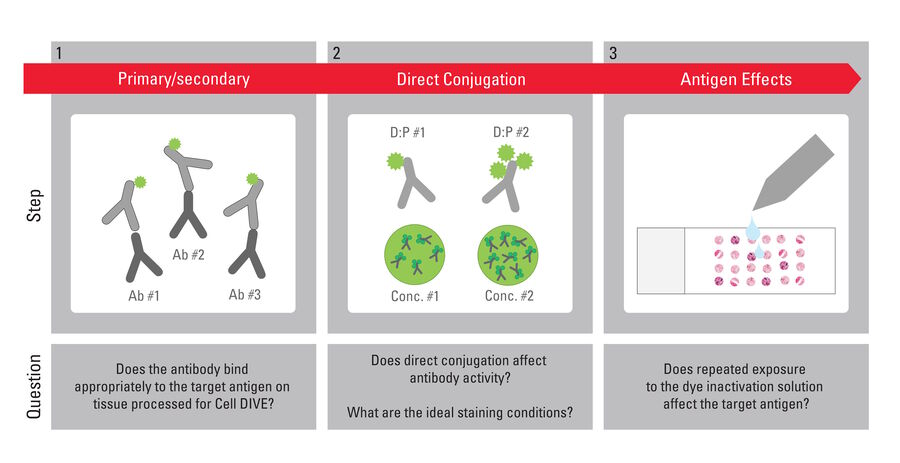
Designed to simplify high-precision multiplexed imaging, the Cell DIVE system includes a catalog of meticulously validated, commercially available antibodies to support a wide range of research and clinical applications. The current list consists of 368 distinct polyclonal or monoclonal antibodies to 308 unique human proteins and 30 antibodies validated for use with mouse tissues. Among them are markers that enable detection of tissue architecture and type, cell segmentation, and determination of cell phenotypes, lineages, and structures. Other validated antibodies reveal biomarkers of drug resistance, immune response, angiogenesis, and a broad range of cancer types, including ovarian, pancreatic, lung, liver, breast, prostate, and endometrial cancer. The rigorous, 3-step antibody characterization process was developed and refined over 10 years in collaboration with academic partners at major cancer institutes. Every Cell DIVE validated antibody has undergone this assessment to guarantee robust and accurate imaging results (Figure 2).
Save time, expenses, and frustration
Performing the comprehensive characterization that each validated Cell DIVE antibody undergoes takes an estimated 32 hours and 1,500–3,000 USD in reagents and supplies per antibody. Both time and expense rise if challenges arise. Thus, a modest study using 20 biomarkers could take up to 16 weeks of work and cost at least 30,000 USD in supplies before imaging of a single experiment begins. Our expert Cell DIVE scientists have done that work for you.
Step 1: Does a commercial antibody meet the specificity and sensitivity standards of Cell DIVE?
For each biomarker, three or more novel polyclonal antibodies or monoclonal antibody clones demonstrated to work on FFPE tissues are selected for evaluation following a thorough review of the literature and life science vendor offerings. The goal of the first evaluative step is to identify antibodies that show (a) preferential binding to the targeted structures with limited non-specific staining, (b) sensitive detection of even low-expression proteins, and (c) staining patterns aligned with publicly available data. References used to confirm experimental outcomes include vendor product profiles, the Human Protein Atlas, and literature reports.
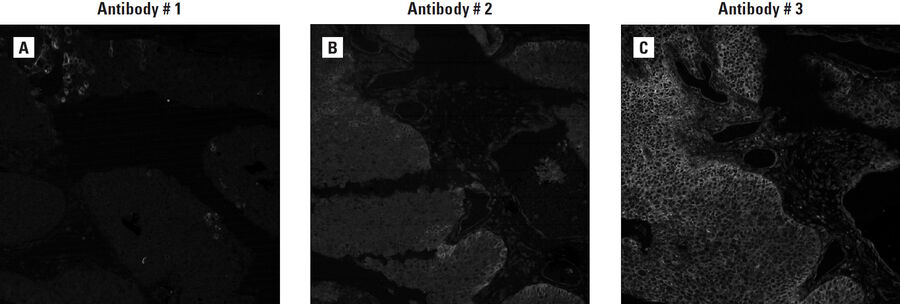
Each antibody is bound to a multi-tissue tumor array of a specific cancer type and then stained with a dye- labeled secondary antibody. An IgG control is included in each evaluation to confirm specificity. Additional controls like blocking peptides, phosphatase treatment for phospho-epitopes, and cell line controls to account for known target expression differences, are used whenever possible. By comparing the staining patterns of each antibody to others (Figure 3), to controls, and to existing data, the top performing one or two candidates are selected for the next characterization step: compatibility with direct conjugation.
Step 2: Does direct conjugation with a dye affect staining performance?
Multiplexed imaging requires creative solutions for immunodetection to avoid cross reactivities common to traditional immunofluorescence with secondary detection. That cross reactivity limits the number of biomarkers that can be explored with the traditional approach. Cell DIVE, in contrast, images multiple markers simultaneously and serially in a single tissue sample. Direct conjugation of dye and antibody avoids the need for secondary detection reagents, creating flexibility in the design of complex multiplexed panels and enabling equalized and efficient use of all available imaging channels. Therefore, the second characterization step aims to determine if direct conjugation alters the staining performance of the selected antibody candidates.
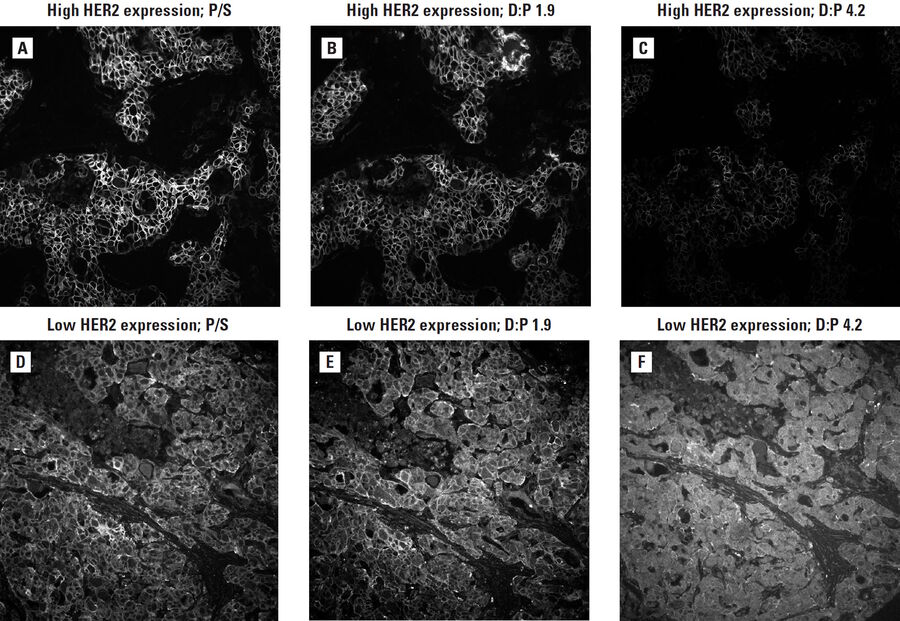
Each antibody is conjugated to a cyanine-based dye at two different dye-protein (antibody) loading ratios and then tested in the same way as step 1 but at a high and a low final antibody concentration. Thus, the staining pattern of the unconjugated primary antibody with secondary detection serves as a benchmark to assess if direct conjugation impacts the sensitivity or specificity of the antibody. Concurrently, the different loading ratios and conjugate concentrations are compared to determine the optimal combination that delivers the greatest signal and appropriate staining with the least background (Figure 4).
Some antibodies show altered staining patterns and reduced signal intensity if direct conjugation interferes with their ability to recognize an antigen. Nevertheless, even when conjugation affects function, the antibody can still be used in the Cell DIVE workflow with primary/secondary detection; it just needs to be used in the first or second staining round to avoid cross reactivity.
Step 3: Does the dye-inactivation process cause antigen effects?
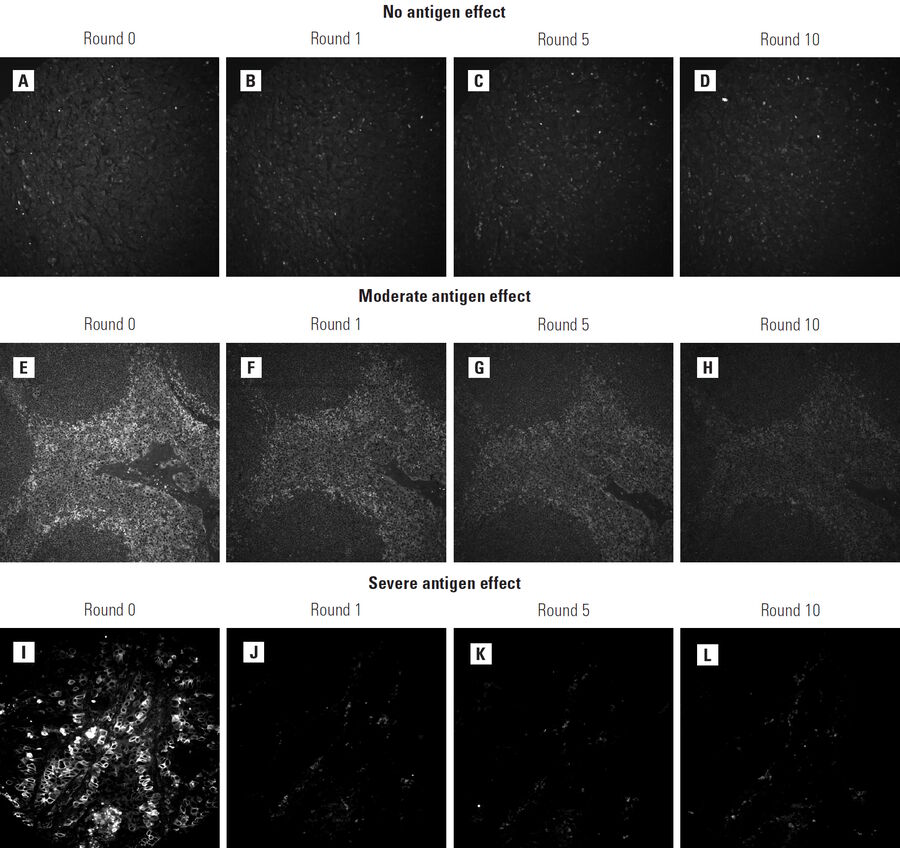
The Cell DIVE dye-inactivation process is generally inert, leaving sample and antibody activity intact. However, a limited number of antigens can be affected by the solution. In such cases, the antibody can still be used in the Cell DIVE workflow but should be imaged in early staining cycles, – before any antigen effects are noted – to ensure that antigen effects do not affect the final results. Therefore, each antibody candidate is tested on tissues that are exposed to multiple rounds of dye-inactivation chemistry. Antibody staining is then compared to an untreated control. A loss of signal intensity or change in specificity between controls and tissues exposed to 1, 5 and 10 rounds of inactivation are likely due to antigen degradation (Figure 5). Initial iterations of the characterization process tested up to 100 rounds of dye-inactivation solution and demonstrated that antigens affected by exposure exhibited effects before round 10.1 The current characterization process was therefore shortened accordingly.
A characterization process designed to make multiplexed imaging easy
The outcome of the Cell DIVE characterization process goes beyond the unequivocal validation of an antibody for the Cell DIVE workflow. Using Cell DIVE validated antibodies also promotes antigen recognition and simplifies study design. The tests are performed using a two-step antigen retrieval protocol designed to unblock a wide range of proteins of interest within FFPE tissues. Therefore, using Cell DIVE validated antibodies also guarantees that this retrieval protocol unmasks all antigens to be tested. Then, each antibody is delivered with the information needed to source it, to define its optimal dye labeling, and to use it under the most effective staining conditions. Cell DIVE validated antibodies support robust study design without the need for iterative optimization.
While the validated list is extensive and growing, an antibody of interest may be missing. In that case, the antibody characterization protocol is included with the Cell DIVE imaging solution. Any antibody can be characterized in a standardized way to craft unique and dedicated multiplexing panels. In this way, Cell DIVE jumpstarts spatial biomarker mapping with ready-to-use tools today, and advances research with the know-how to expand analyses tomorrow.