Introduction
Genetics seeks to understand how genes determine an organism's phenotype. Forward genetic screening has revolutionized this pursuit by generating unbiased genotype-phenotype maps. Through the creation of genetic mutant libraries and the subsequent isolation of interesting phenotypic variants for genotyping, researchers discover connections between genes and their resulting traits in an unbiased manner. Accelerated by techniques for targeted gene disruption, this methodology now scales to genome-wide screening. So far, these screens have been limited to investigating simple phenotypes that inherently cause clonal enrichment or result in changes in fluorescence intensity measured by FACS.
Increasingly powerful microscopic imaging provides information-rich data on diverse cellular phenotypes. When coupled with recent advances in deep learning, this presents an ideal technology to read out biological phenotypes of interest in genetic screens. However, due to technological constraints it has up till now found limited application.
What can you expect in the webinar?
Sophia Mädler will present spatially resolved CRISPR screening (SPARCS), a platform for microscopy-based genetic screening for spatial subcellular phenotypes at the human genome scale. It uses automated high-speed laser microdissection to achieve isolation of phenotypic variants in situ from virtually unlimited library sizes. Using deep learning, phenotypically interesting cells can be identified on the basis of single-cell images.
The power and potential of SPARCS
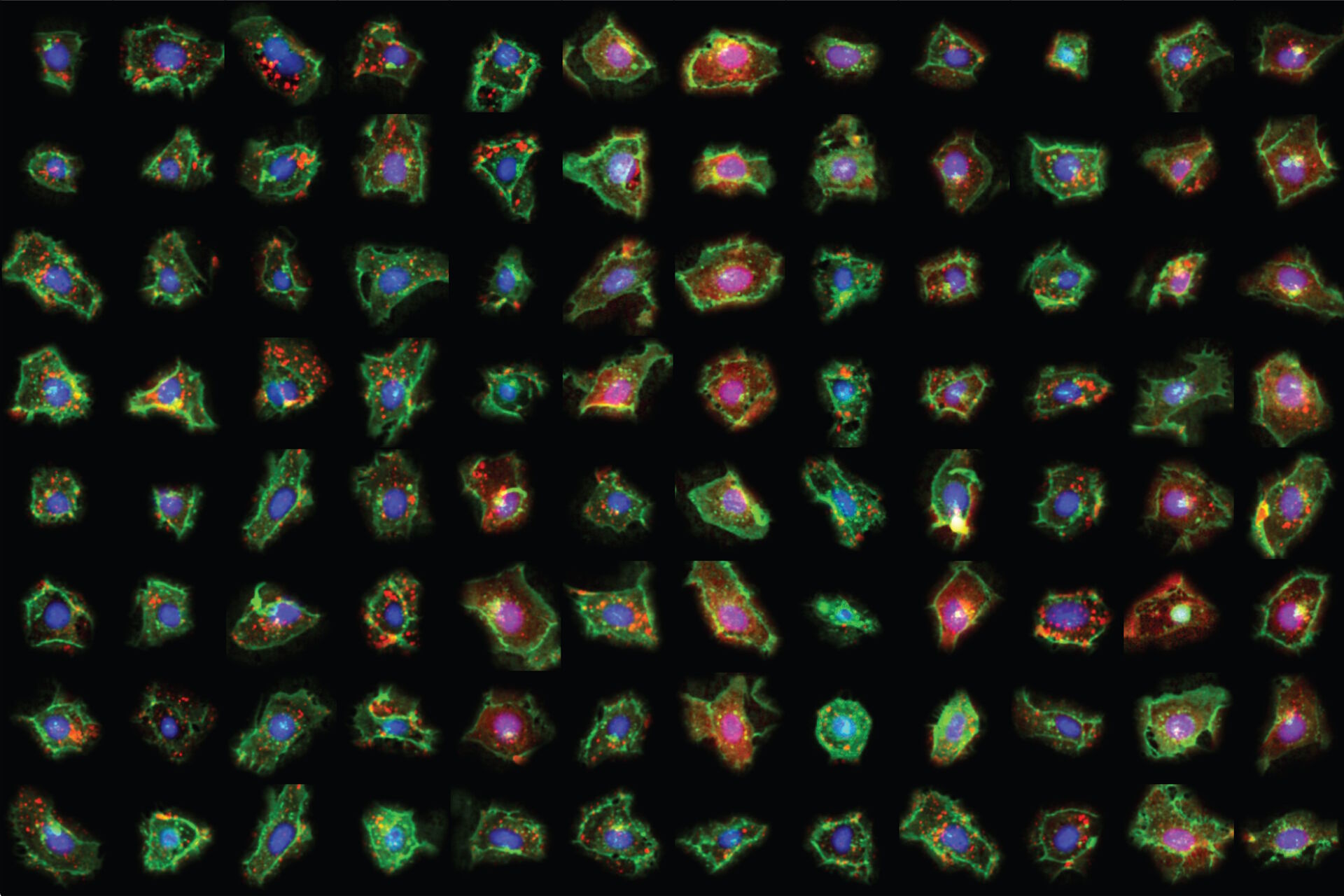
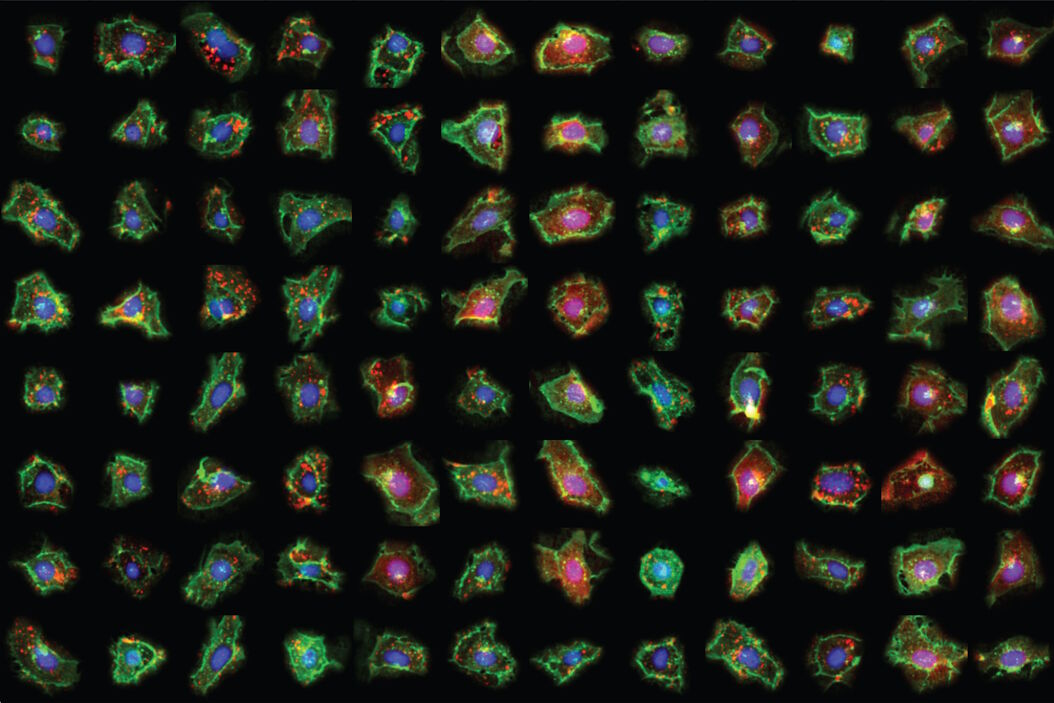
To demonstrate the potential of SPARCS, Sophia will present the results from a genome-wide CRISPR-KO screen on autophagosome formation in 40 million human cells.
Phenotypic variants showing defects in autophagosome formation were identified using a convolutional neural network (CNN) classifier which recognizes differences in sub-cellular distribution of the autophagosome marker LC3B based on single-cell images. Using this classifier, SPARCS recovered almost all known macroautophagy genes in a single experiment and discovered a role for the ER-resident protein EI24 in autophagosome biogenesis.
With its focus on open and accessible design, SPARCS can be seamlessly integrated with any microscope. Coupled to newly developed open-source python libraries, SPARCS allows for the rapid generation and analysis of single-cell images. Using state-of-the-art deep learning methods, cells can be characterized based on their subcellular spatial phenotypes. Identified cells of interest can then be rapidly translated into a cutting map for laser microdissection.
Throughout the webinar, Sophia will outline how SPARCS and its developed tools can be used to harness the full power of advanced imaging technologies to enable genome-wide forward genetic screening for diverse spatial sub-cellular phenotypes in situ.