Time-lapse microscopy
Time-lapse microscopy [1,2] involves capturing images of a specimen via a microscope at a specific rate, usually mentioned as frames per second (fps), over a certain period of time. If the image-capture rate is lower than the viewing rate used to observe the recorded images, then time appears to be moving faster when observing the sequence of images like a video. Similarly, if the image-capture rate is higher than the viewing rate, then time seems to be moving slower. Therefore, time-lapse microscopy enables the observation of long microscale events, i.e., which occur over minutes, hours, or days, to be viewed within seconds, minutes, or hours. Time-lapse microscopy is used for the study of live specimens, such as cell cultures, acute tissue samples, and model organisms as they grow and develop over time, to gain more insights about biological processes [3,4]. For instance, cell migration is important for multicellular organisms during embryonic development, tissue repair, functioning of the immune system, and tumor invasion. In order to accurately follow the changes of cells and organisms over time, then “longitudinal” studies [5] are used where the same sample or specimen is observed under specific conditions over a defined period of time.
Spheroids
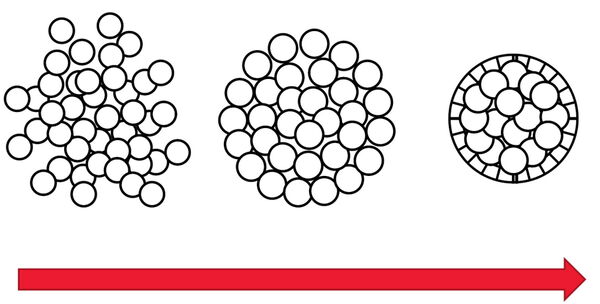
Spheroids are a type of 3D cell culture, like organoids, which mimic the physiological functions of living tissues and organs [6]. Monolayer 2D cell cultures are normally cells simply growing flat on a substrate. Spheroids and other 3D cell cultures can grow into large volumes that enable 3-dimensional cell-cell interactions which is more like native tissue in an organism. These 3D cell cultures can be used for research to help obtain a better understanding of cells in a more real-life microenvironment. Spheroids are useful for applications in neuroscience, regenerative medicine, as well as cancer and cardiovascular research.
Challenges
When doing time-lapse microscopy experiments with spheroids, there are certain challenges which can arise. As the experiments can last for several days or even weeks, prolonged sample survival must be achieved which requires that near physiological conditions are ensured. Expression of fluorescent markers must be kept at endogenous levels to prevent impairing cell homeostasis. There must be a stable supply of nutrients and an unchanged concentration in the medium which can be impaired by evaporation. Proper imaging over long times requires that microscope imaging stay in focus and that it is adapted to changing sample characteristics, e.g., lateral and axial growth.
Research labs may not have the instrumentation needed for time-lapse microscopy work involving 3D cell cultures. In this case, normally researchers must have access to shared facilities utilized by multiple users to get around this problem, but that can mean long waiting times before the needed instruments are available. These challenges can lead to delays in acquiring the important, quantifiable results that lead to fundamental breakthroughs and the gaining of new insights.
Introducing Mica
Mica, the world's first imaging Microhub, unifies everything a researcher needs in one completely controlled, highly flexible environment, supercharging microscopy workflows to get meaningful scientific results faster. With Mica, users can benefit from:
- Access for all: Easy setting up of controlled environmental conditions, matching focus strategy, and setting up the imaging conditions
- Radically simplified workflows: On screen annotation and extraction of multiple parameters
- No constraints: Optimal environmental conditions and multiple imaging modalities (brightfield, widefield, and confocal) to match experimental needs and intelligent automation for water immersion and focus strategies
Methods
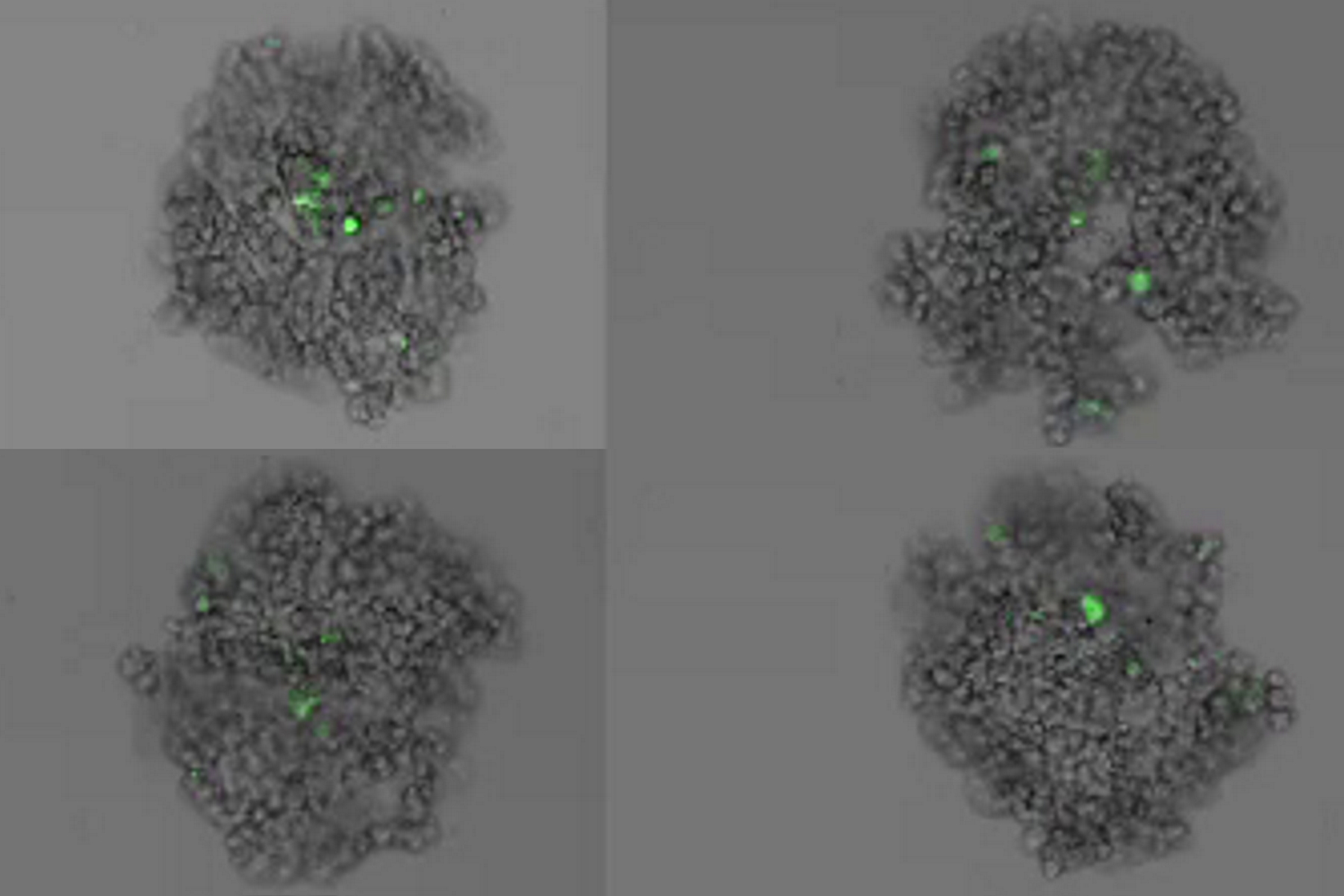
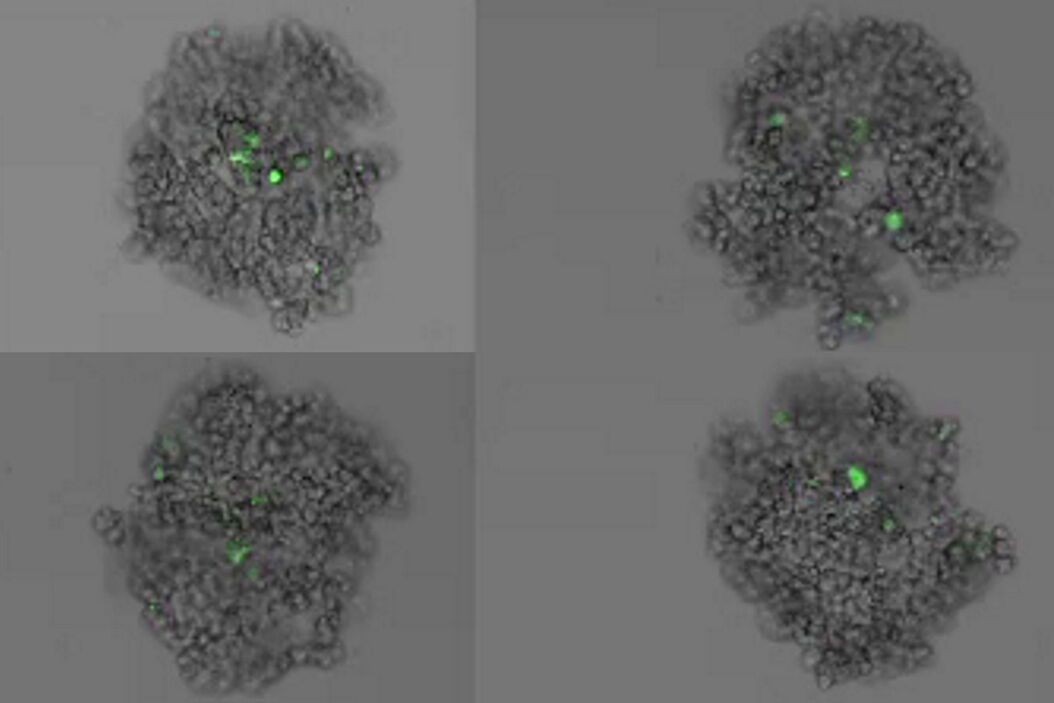
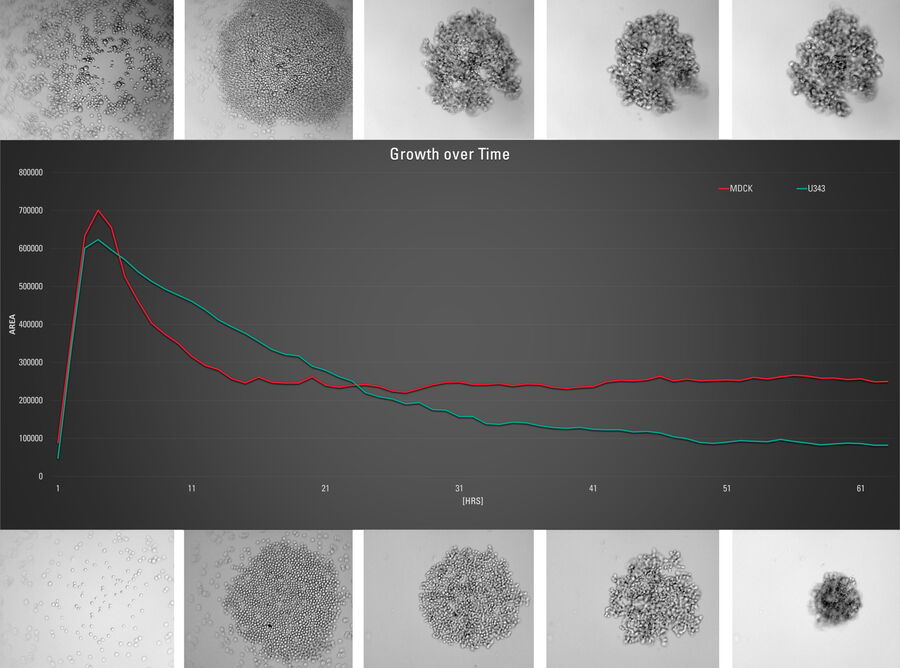
This long-term time-lapse study involves the use of Mica to investigate the formation of spheroids starting from either U343 cells or MDCK cells stably transfected with MX1-GFP. Growing spheroids requires optimal physiological conditions which ensure undisturbed cell cycles and proliferation [6,7]. Previous results show that the growth itself is often correlated with the expression of specific proteins or markers for certain hallmarks of cell-states and differentiation [6,7].
Mica is used for this key application as an incubator to maintain 3D cell cultures and spheroids under near physiological conditions and with minimized media evaporation. Mica allows users to measure spheroid growth and to analyze protein-expression levels.
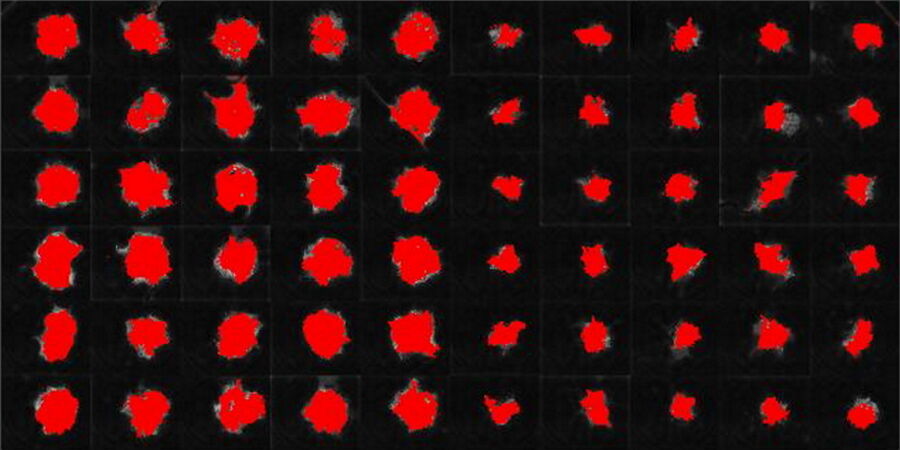
Images and data acquired during the long-term time-lapse study of the formation of 3D spheroids and their growth are shown in figures 2 and 3.