Cell Culture on EM grids
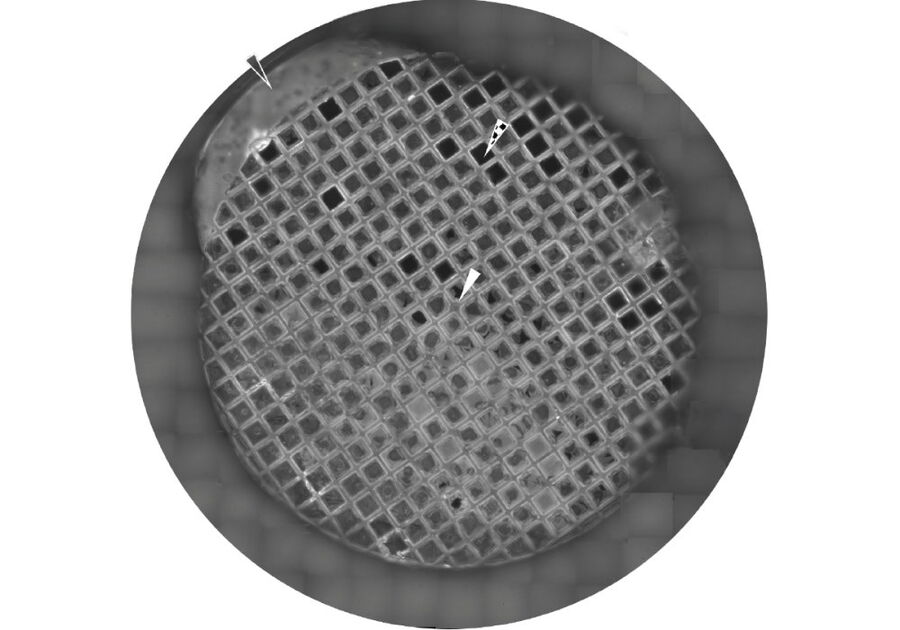
Typically, acutely isolated or cultured cells are seeded on gold or titanium grids with a perforated carbon film (e.g. QuantifoilR) or an SiO2 film (Fig.1, Mahamid et al. 2019). Titanium and SiO2 seem to be stiffer and more stable for the subsequent steps and no additional carbon layer is needed (Toro-Nahuelpan 2019).
Grids are bioactivated by poly-L-Lysin or Fibronectin, trypsinated cells are seeded over night to allow attachment to the carbon layer for the following steps (Mahamid et al., 2019).
Micropatterning
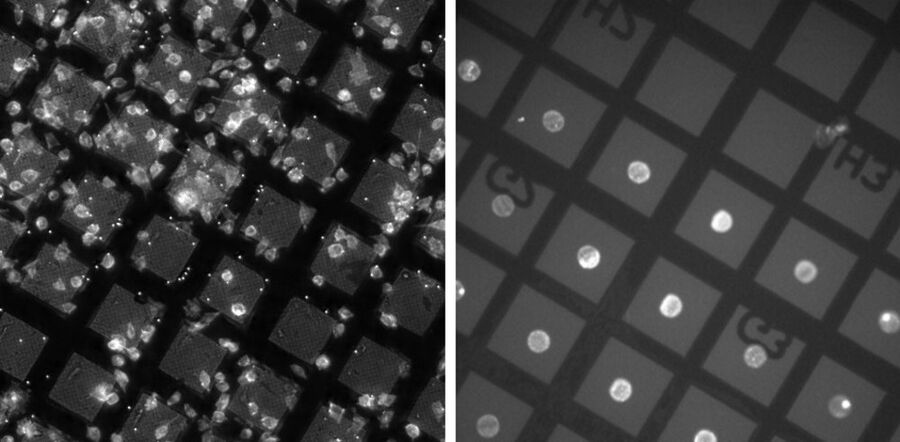
For access to cellular samples for successful FIB milling and subsequent analysis in the cryo-TEM, it must be ensured, that the cells are located in or close to the center of the grid squares. Unfortunately, cells tend to grow on grid bars or grow in clusters and are therefore not suitable for FIB milling and analysis with electron transmission. To overcome this challenge, micropatterning allows users to control the location and spreading of cells on the carbon film (Fig. 2), increasing the reliability of the related workflows.
The surface of the grids is coated with polyethylene glycol (PEG) which prevents the attachment of biological material. By removing the coating with a UV laser, the adherence of cells can be specifically controlled and the accessibility for FIB milling and TEM ensured (Toro-Nahuelpan 2019). In addition, specific patterns can be created, thereby influencing the complete cellular architecture and facilitating the investigation of biomechanical phenomena with cryo-electron microscopy.
visualizing F-Actin). Right image: Precisely positioned cells in the center of grid squares made accessible for the FIB (fibroblasts adhering on fibrinogen micropatterns; images kindly provided by Alvéole in collaboration with Prof. Dr. Kay Grünewald, CSSB, Hamburg, Germany.
Plunge Freezing
To keep the samples as close to the native state while being fixed for the electron microscopy, the cells must be very quickly frozen without the creation of destructive ice crystals. This process is called vitrification as the ice becomes amorphous and glass-like (vitreous).
To achieve this for the sample cells, the grids must be rapidly plunged into a suitable cryo-gen (typically ethane, or ethane-propane). In 1981, Jacques Dubochet published the first manual blot-and-plunge method that is still widely used to reach excellent results (Dubochet, J. & McDowall, A. W., 1981).
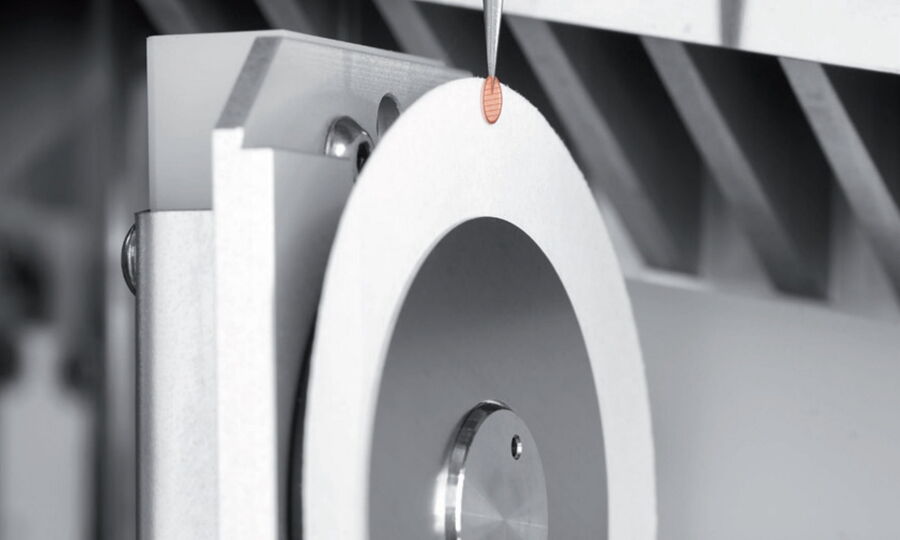
Before the plunging, excessive liquid must be removed. The standard technique is controlled blotting with filter paper (Fig. 3, Dubochet, J. et al. 1982; Bellare et al., 1988; Frederik, P. M. et al. 1989).
There are different commercial devices available such as the EM GP2 for automatic blotting and plunge freezing. According to the various needs of different sample types, several protocols for sample preparation involving blotting are available (see also here).
Storage, loading and transfer under cryo-genic conditions
After the vitrification the samples must be maintained under cryo-genic conditions throughout the complete workflow. Hence all steps from storage through transfer to the different imaging systems must be performed so that the sample is not devitrified and/or contaminated. This is particularly difficult as such deep cold samples act as a trap for humidity and dust in the close vicinity. Researchers and manufacturers made extensive efforts to develop and provide solutions for keeping the samples safe throughout the different steps of the workflow.
Samples are generally stored in grid boxes in a group of four and the grid boxes are maintained in perforated Falcon tubes in larger LN2 tanks. There are also more sophisticated puck systems available.
For transfer and loading into sample holders, LN2 is typically used. Unfortunately, LN2 tends to host crystalline ice contaminations derived from the humidity in the air after a while. The ice tends to attach to grids while being transferred, disturbing the subsequent milling and imaging. In addition, the visibility inside of the LN2 is poor as it is constantly moving, and streaks are persistent.
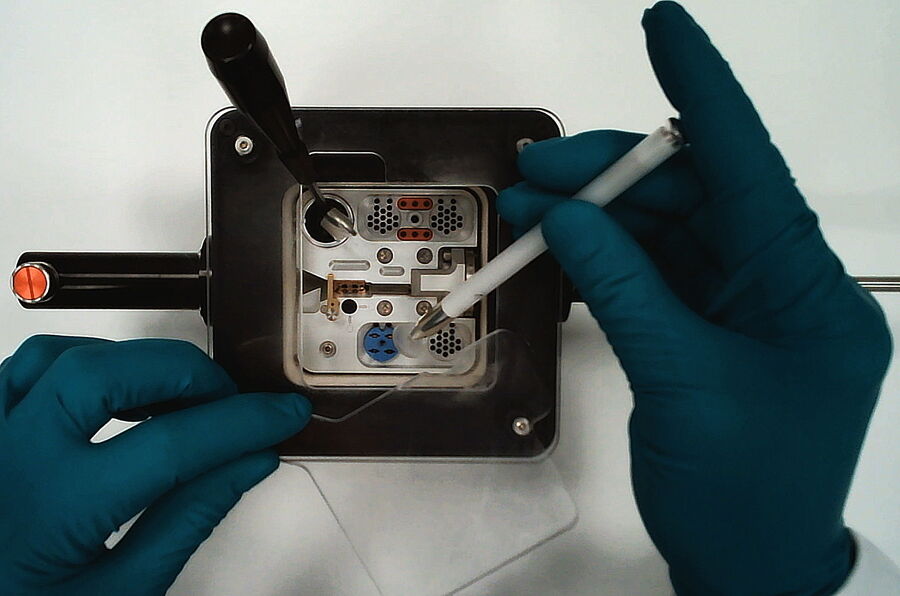
Therefore, it is beneficial to load and transfer the sample within the gas phase above the LN2 keeping safe cryo-genic conditions while providing excellent visibility for the loading (Fig. 4).
Leica Microsystems has experience with loading and transfer equipment under GN2 for more than 30 years. The new Cryo-Microscopy Kit was developed based on this experience, also implementing the feedback of many customers into a state-of-the-art shuttle and cartridge system.
Check of quality and target distribution
As cryo-workflows in general, but EM operation time in particular are quite expensive, an early quality check of the sample is crucial. Many factors contribute to the decision if a sample can be transferred to the next workflow step, i.e., the structural integrity of the carbon foil, the vitrification quality including the ice thickness and its distribution, the existence, distribution and accessibility of the cells-of-interest and finally the presence and localization of the structure of interest.
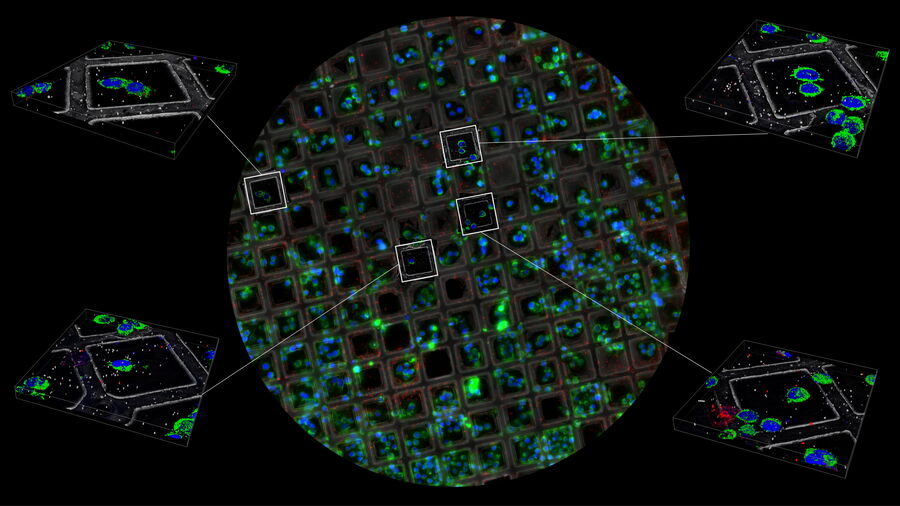
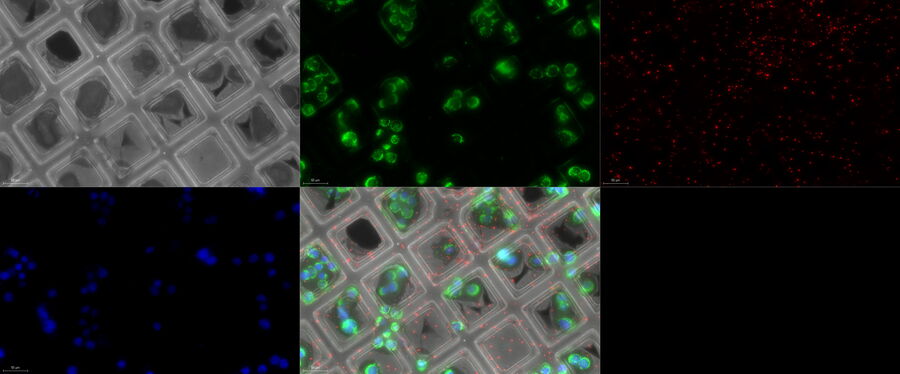
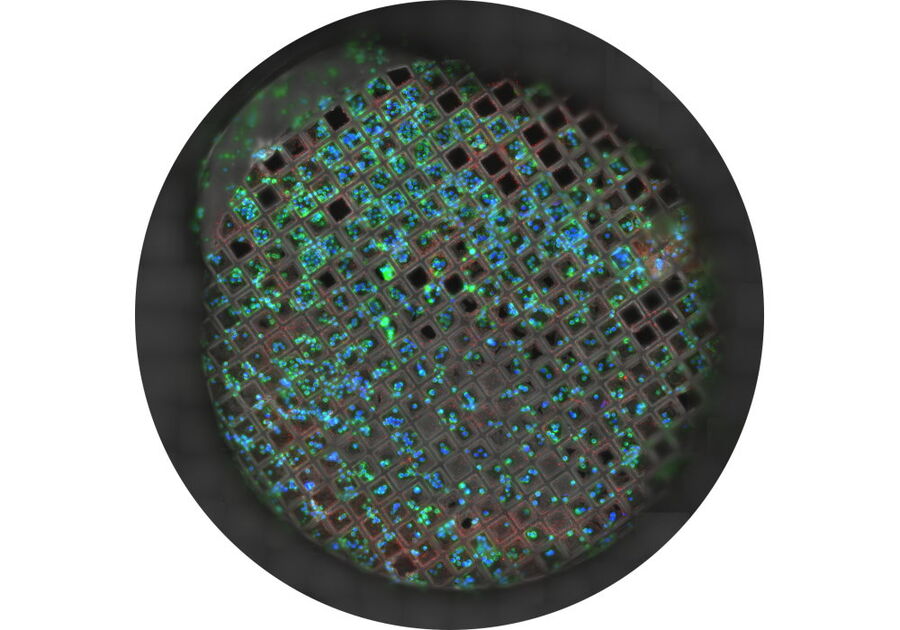
All these parameters can be checked by a camera-based cryo-light microscope such as the THUNDER Imager EM Cryo-CLEM or by using the camera mode on the STELLARIS cryo-confocal microscope (Fig. 5).
The transmission mode reveals the grid, foil and cell quality, the reflection image shows the surface of the grid and reveals in particular the vitrification quality and ice thickness, while the fluorescence image provides information about the expression level of distinct target proteins and their distribution.
Within the LAS X Coral Cryo-software workflow, the users are guided to automatically create in-focus, crisp overviews of the complete grid with different image modalities.
Mark landmarks, lamellae points and bead centers
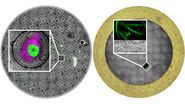
To correlate 3D images of the cryo-LM and the subsequent cryo-FIB-SEM/TEM images, first an overview image of the grid is needed to roughly align the two image modalities (Fig. 6). Here, the reflection images are helpful as they are similar to the SEM images, but also transmitted light mode can be used. The center mark and other landmarks such as defects in the carbon layer allow to quickly orient and align the overviews.
Second, the super-resolved confocal 3D images are needed. These image stacks are used to perform a high-accuracy correlation within the volume of the potential lamella position. After the overview alignment, it is possible to find the correct positions of the 3D confocal stacks for subsequent high accuracy correlation. As a precondition, the image positions relative to the overview and to each other must be provided. This is the case when following the Coral Cryo-software workflow (Fig. 7).
The set of both the camera overview and super-resolved 3D images is essentially needed to retrieve the target positions and to define the milling position on the FIB-SEM. This is particularly important, as in a standard FIB-SEM the fluorescence and therefore the target site cannot be made visible.
Recent developments of EM manufacturers provide FIB-SEM-integrated fluorescence microscopes, an excellent option to increase the reliability and accuracy of the workflow by checking the fluorescence during the milling process. However, these systems do not have the necessary resolution and flexibility of acquisition modalities to enable precise 3D targeting as with a stand-alone cryo-confocal system.
How to correlate and retrieve lamellae positions
As a commonly used minimal standard, researchers use a screenshot of the LM images to retrieve the target xy-coordinates on the EM. Unfortunately, the side-by-side comparison of images is time-consuming and prone to failure and therefore not reliable. As a workflow provider, Leica Microsystems sought to improve this situation on its THUNDER Imager EM Cryo-CLEM. Landmarks and target markers can be positioned on the images and exported as complete sets in open EM formats. As a first step, this was implemented for 2D, yet the next logical step was to improve the resolution and to expand the coordinate system towards 3D.
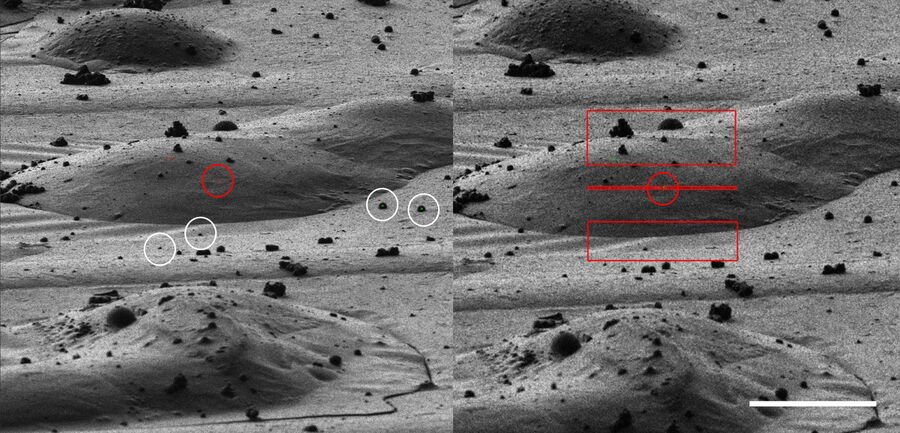
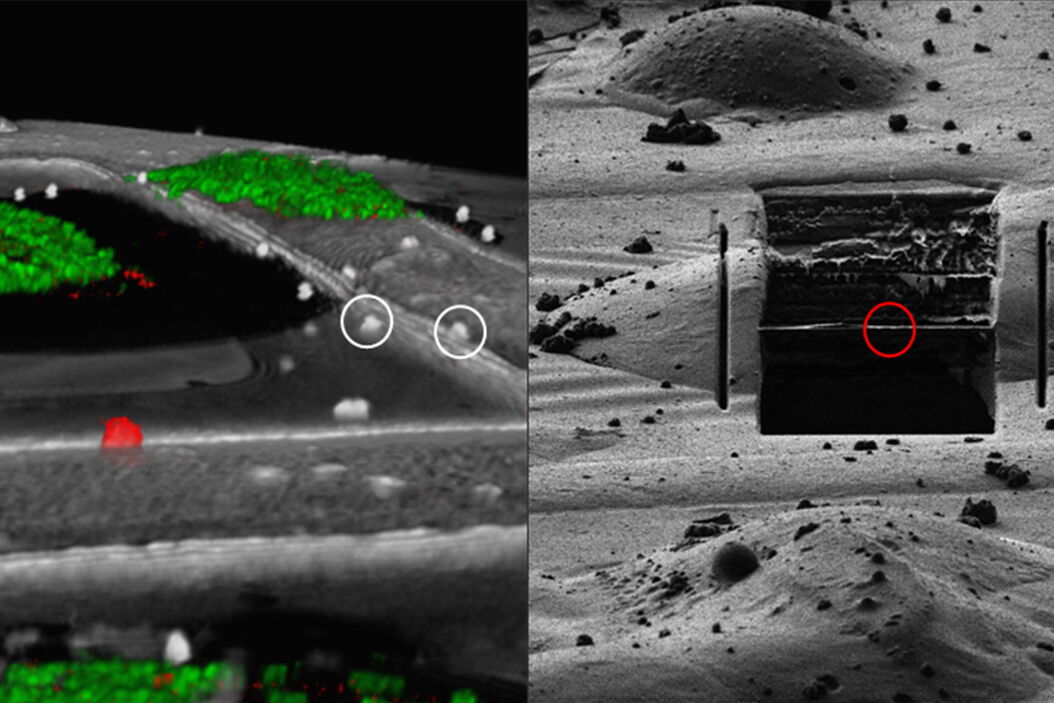
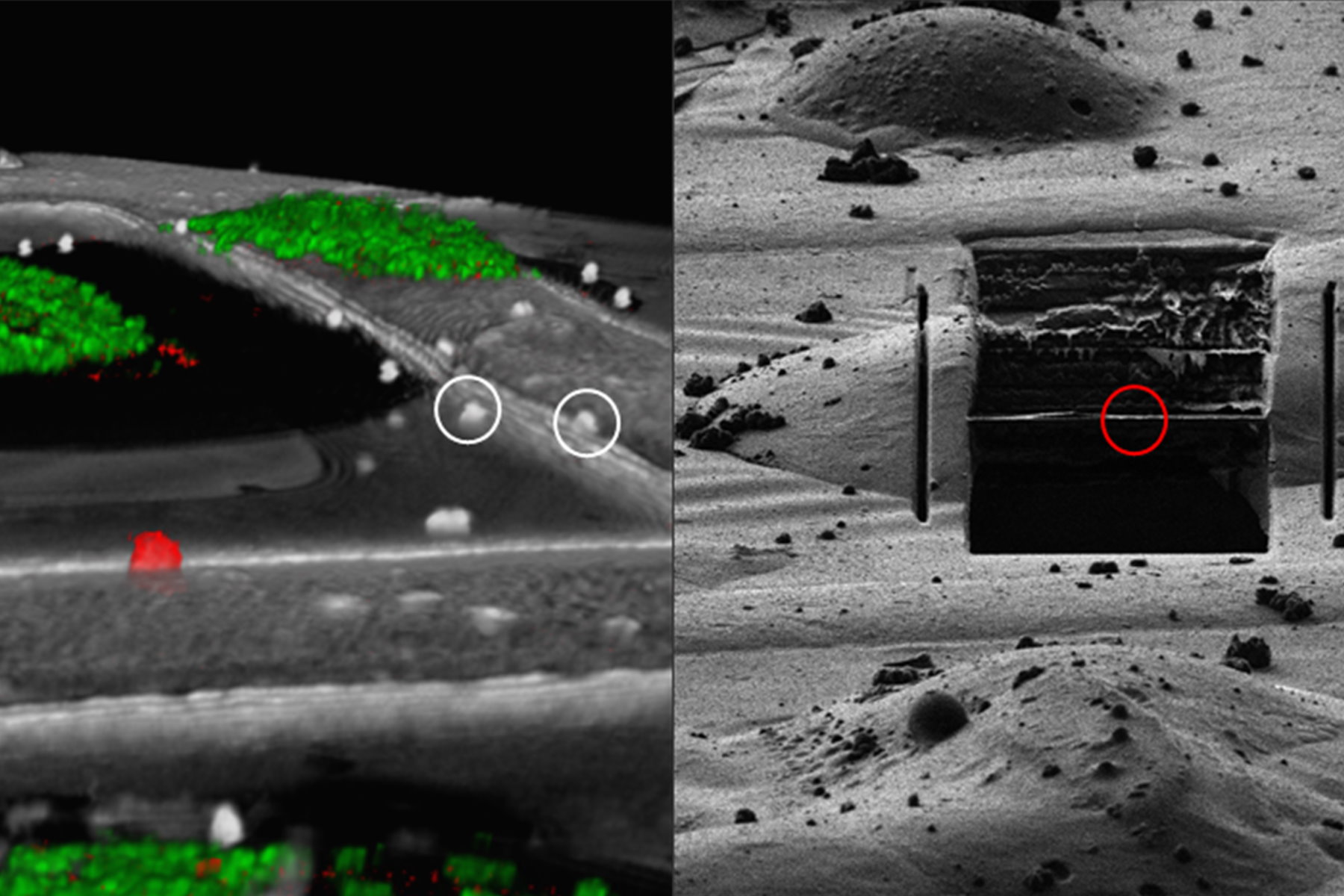
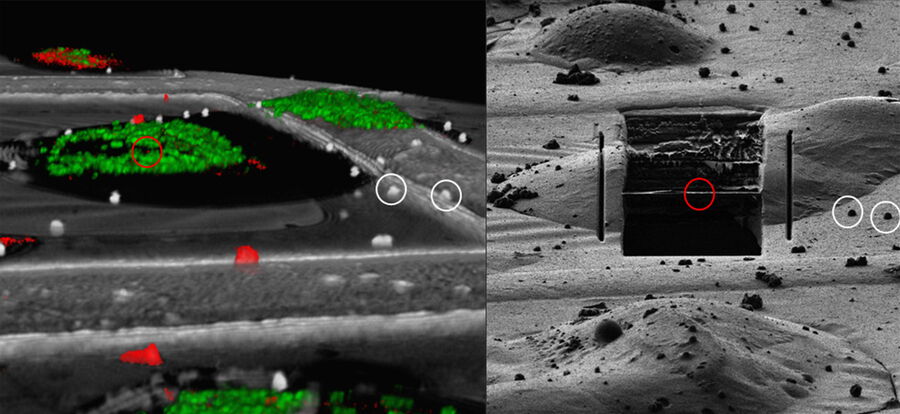
For high precision correlation and 3D targeting a bead-based method has become widely used (Alegretti et al., 2020; Klumpe et al. 2021; Bieber, A., Capitanio, C. et al. 2021).The beads are mostly added to the cells prior to vitrification and are visible in both the LM and EM and serve to align the stack in xyz as basis for the correlation of the image data and hence the proper positioning of the FIB milling windows (Fig. 8).
Typical beads are 1 μm in size and completely spherical which enables sub-diffraction fitting of their center coordinate. Microbeads that contain metals can be more clearly visualized via back-scattered electrons in the SEM, distinguishing them from similar size ice crystals. Beads are preferentially chosen, such that their fluorescence emission is different from that of the actual target, to enable better discrimination.
To use the 3D data from the cryo-LM and the FIB SEM for cryo--LM guided lamella preparation, there is an open source software available (3D correlation toolbox, 3DCT, Jan Arnold et al., 2016).
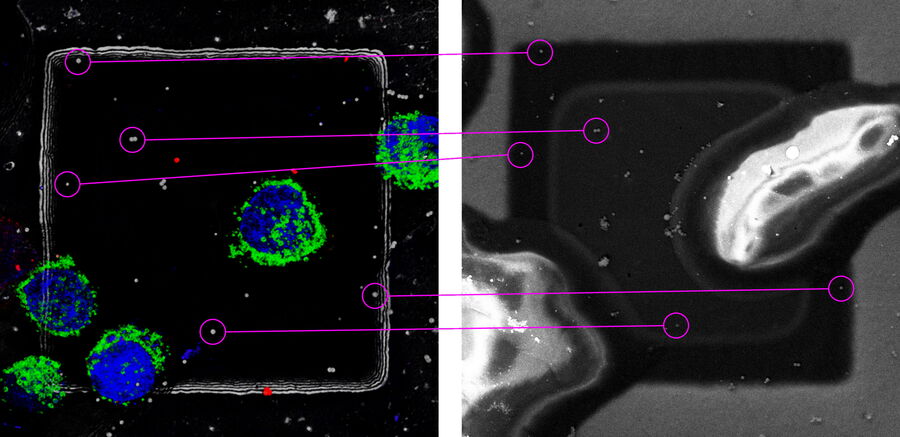
The cryo-LM images are loaded into the software running on the FIB-SEM. A three-point correlation between the 2D LM overview and the SEM images is used for the initial orientation. Afterwards, the relevant field of view is taken with the ion beam and the identical beads in the LM stacks and in the FIB image are clicked manually. In Figure 10, an LM and a FIB image is shown with a target site and beads resembling the arrangement in a targeting software.
The software determines the center of the beads very precisely by a Gaussian fit of the signal in x, y and z. Recent implementations add a semiautomatic bead detection and other features for an easier performance of cryo--FIB workflows. (SerialFIB, Klumpe et al. 2021).
Several beads on the grid bars surrounding the final target structure are selected and serve as coordinate system for the milling. The underlying calculation is a linear-affine transformation considering scaling, rotation and translation. Finally, the target structure is selected in the LM image and superimposed to the FIB image.
According to the position of the target structure, the milling windows can then be positioned (Fig. 10).
How do I benefit from the Coral Cryo-workflow?
The Coral Cryo-software workflow is developed to feed the bead-based targeting workflow. It provides the imaging jobs needed to create an in-focus camera overview image (Fig. 6 and 7). All necessary autofocus functions can be adjusted and assigned properly, and potential lamellae positions can be marked, and super-resolved confocal z-stacks performed at the defined positions.
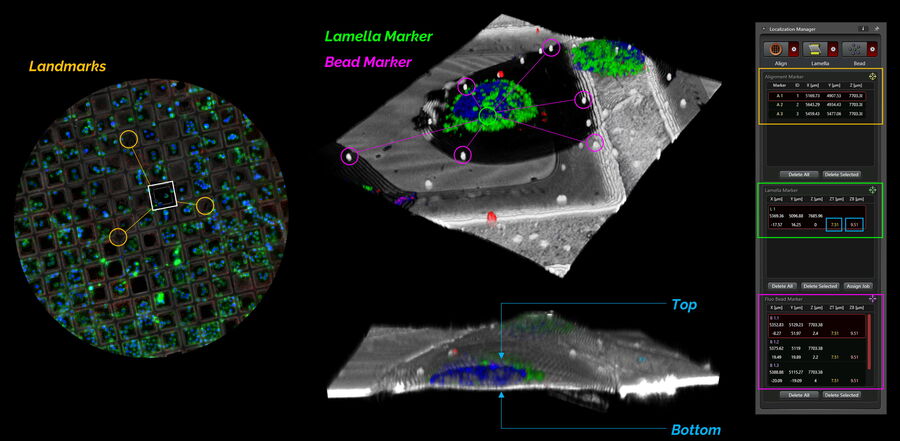
Within the Localization Manager (Fig. 11), all necessary coordinate markers can be determined and are provided in an open format (*.xml). The images are saved automatically in a data format suitable to be implemented into any FIB-SEM software (e.g., tif, ome.tif, imageJ-tif…).
Alignment markers serve to perform the initial rough alignment based on landmarks using the camera overview. Lamella markers serve a two-fold purpose: as position markers for performing a super-resolved confocal 3D scan and as precise 3D marker for the target structure(s) after the image acquisition. The subpixel interpolation ensures high precision targeting within the 3D image at this stage. Finally, the interpolation is also used to mark the bead coordinates for the subsequent bead correlation on the FIB-SEM.
Cryo-FIB Milling
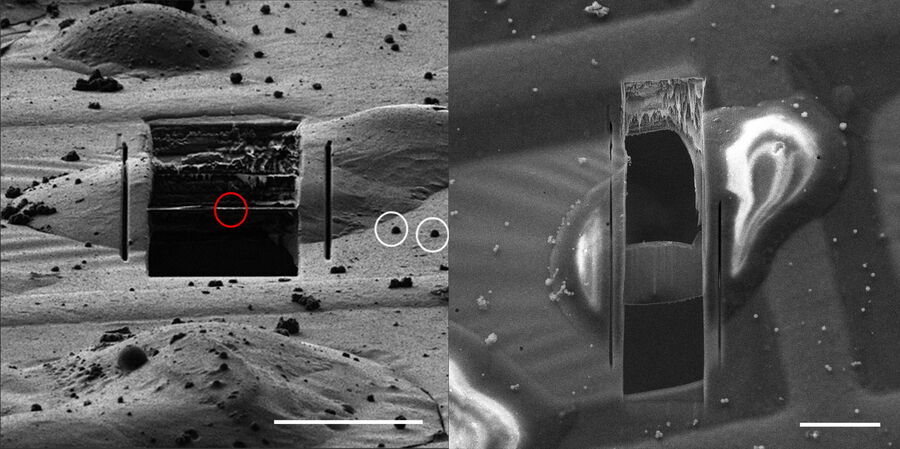
After the necessary correlation is performed and the milling windows are set, the lamellae sites are usually roughly milled down to approx. 1 μm and afterwards final polishing steps towards electron transparency are performed (Fig. 12).
The reason for a two-step approach is that ice contamination and/or milled material might be deposited on the lamellae. In order to avoid ice condensation on the final lamellae, a fast-polishing process is recommended (Schaffer M et al. 2017). Automated approaches with open source and commercial software are available for the milling as well.
Cryo-Transmission Electron Microscopy
After cryo--FIB milling, the grid with the thin lamellae is transferred to the cryo--TEM, where a tomographic series of images is acquired as the grid and hence the lamellae are tilted incrementally. The images are processed computationally to reconstruct a 3D tomogram of the recorded volume. By averaging many images of the sample, the inherent noise can be reduced so that higher resolution structures of particles, i.e., proteins or protein complexes, can be obtained. This process is called sub-tomogram averaging (Wan & Briggs, 2016; Zhang 2019). Conceptually this compares to single particle analysis, yielding down to sub-nanometer resolution of macromolecules in situ.
Summary
In this article we show that cryo-confocal microscopy is an essential component in cryo-workflows to assess the quality and target distribution of vitrified samples on EM grids. The highly resolved confocal data recorded under cryo-conditions enable the identification of structures of interest under fluorescence in 3D. Furthermore, the 3D volume serves as reference for the correlative approach to retrieve the target structures in the FIB-SEM for milling and subsequent electron tomography in the cryo-TEM to achieve sub-nanometer resolution of the targets.
The Coral Cryo-workflow with the new confocal platform STELLARIS and the Coral Cryo-software enables even unexperienced users to create grid overviews, super-resolved 3D images and precisely defined coordinate markers to ensure a solid basis for the subsequent FIB milling and cryoelectron tomography.