Assessing RNA quality of differently prepared murine-brain tissue sections
Due to its instability, RNA is generally more difficult to work with than DNA. RNA does not have the stability of the DNA double helix. To obtain the good results when dealing with RNA it is essential to start out with high-quality material and to carry out particularly careful quality control before and after processing. The so-called RNA Integrity Number (RIN) is an indicator of the RNA quality [1]. On a scale of 1–10, a RIN value of 1 indicates that the RNA is completely degraded and a RIN value of 10 that the RNA is fully intact. The higher the RIN number, the better the RNA quality. Wherever possible, material with high RIN numbers should always be used.
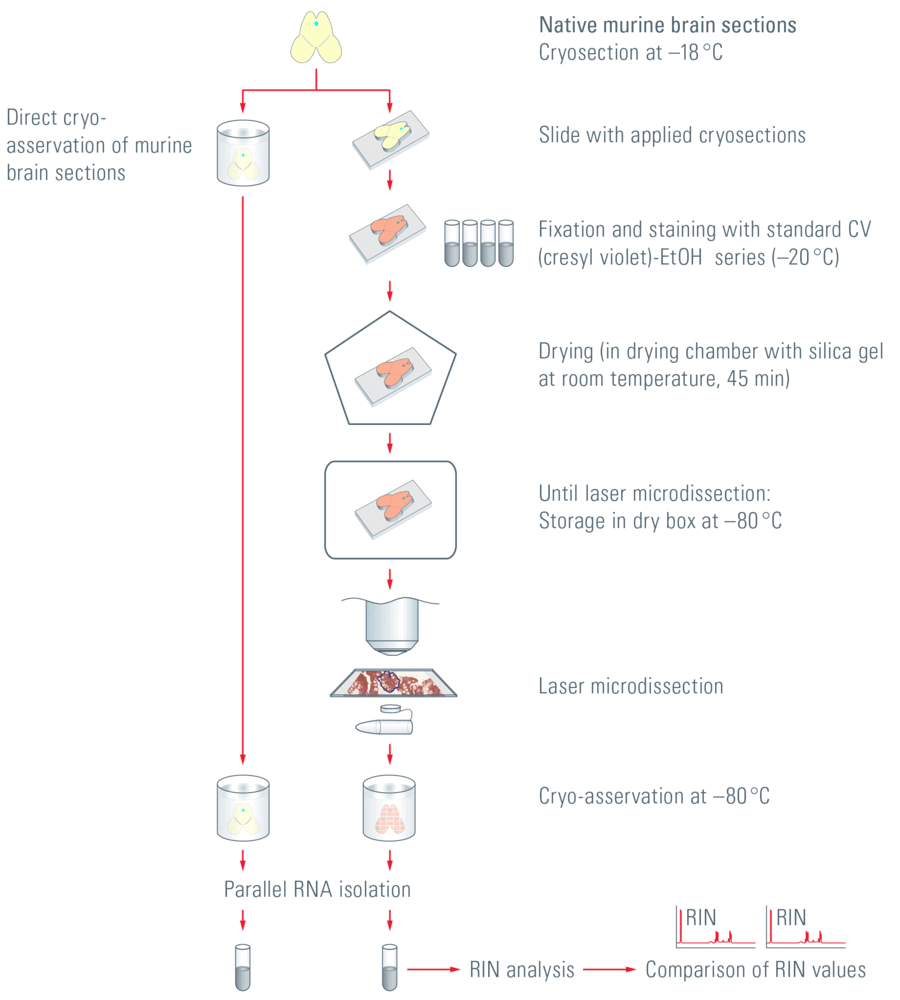
The influence of preparatory fixing and staining protocols and the subsequent UV laser microdissection of murine-brain tissue sections (postmortem) on the RNA quality is described below. The RIN numbers of cryo-preserved, untreated brain sections were compared with sections that were treated and isolated according to standard protocol. Figure 1 shows the sample preparation workflow from cryosection to RIN analysis.
Cryosection with subsequent staining and fixing
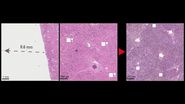
To compare untreated tissue samples with fixed and stained tissue samples, 3 cryo-preserved murine brains of known RNA integrity (RIN) were used. After cryo-preservation at –80° C, these were transferred to a cryo-microtome and incubated at –35 °C for 45 min. Then the tissue was equilibrated at –18° C for 30 min and cut into 12 µm sections at –18° C by cryo-sectioning. The murine-brain sections were divided into two groups which were treated in the following ways. One, the control group, was directly frozen at –80° C. The other group was applied to an RNase-free, UV-treated PEN slide. This was done by briefly thawing the sections at room temperature and fixing them for 2 min in 75% ethanol cooled to –20° C. After fixation, they were immediately stained with drops of sterile-filtered cresyl violet (CV) dye solution (5% cresyl violet in pure ethanol) and incubated for 45 secs with a slight to-and-fro movement. After this staining procedure, the dye solution was briefly drained off and the slide was immersed in a sequence of ethanol solutions with increasing concentrations (75%, 95%, and 100%) for 4 secs each and then finally fixed in 100% ethanol (test group). Following this fixing and staining procedure, the slide was placed in a drying chamber with silica gel for 45 mins at room temperature and then stored at –80° C in a dry box pending laser microdissection.
UV laser microdissection
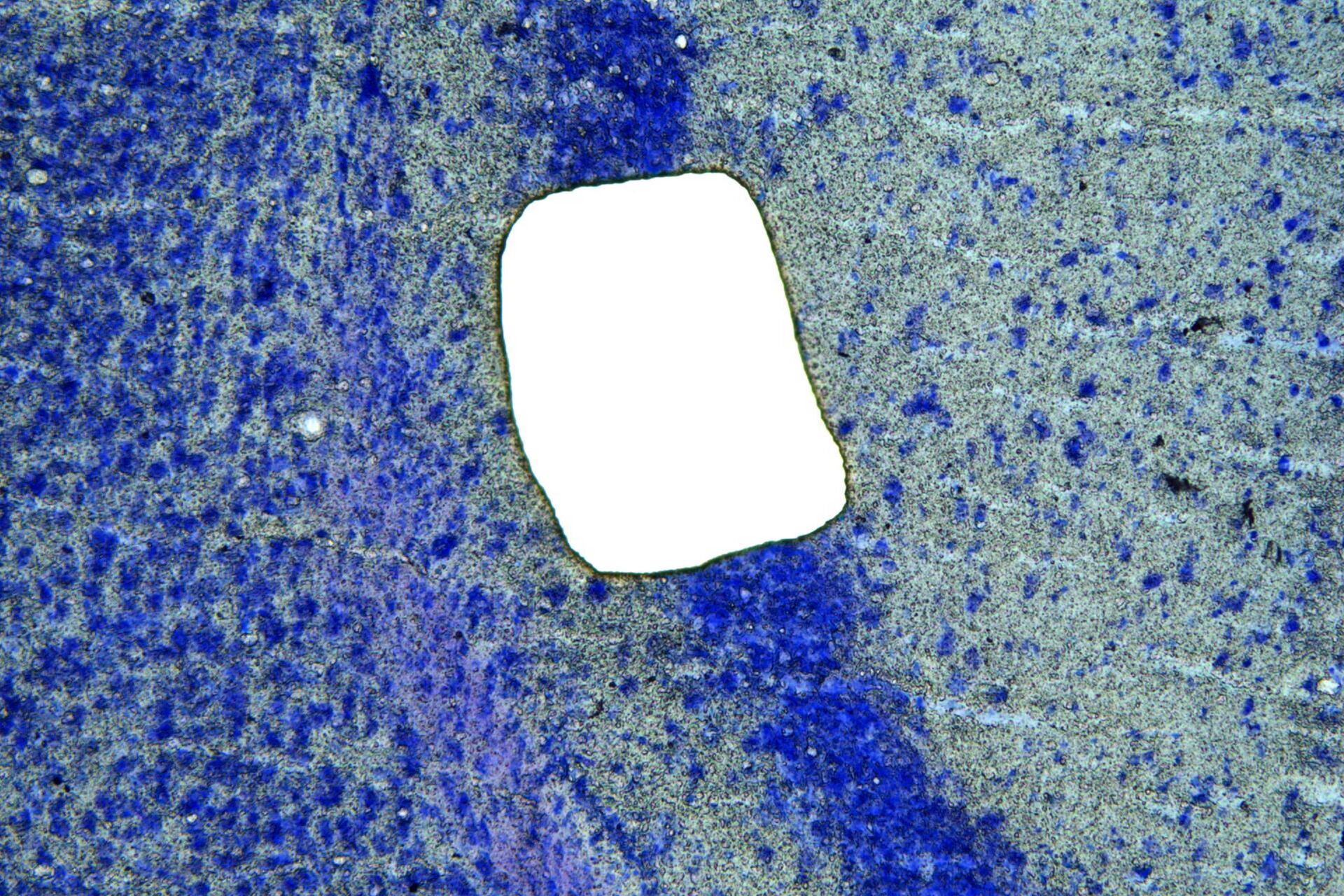
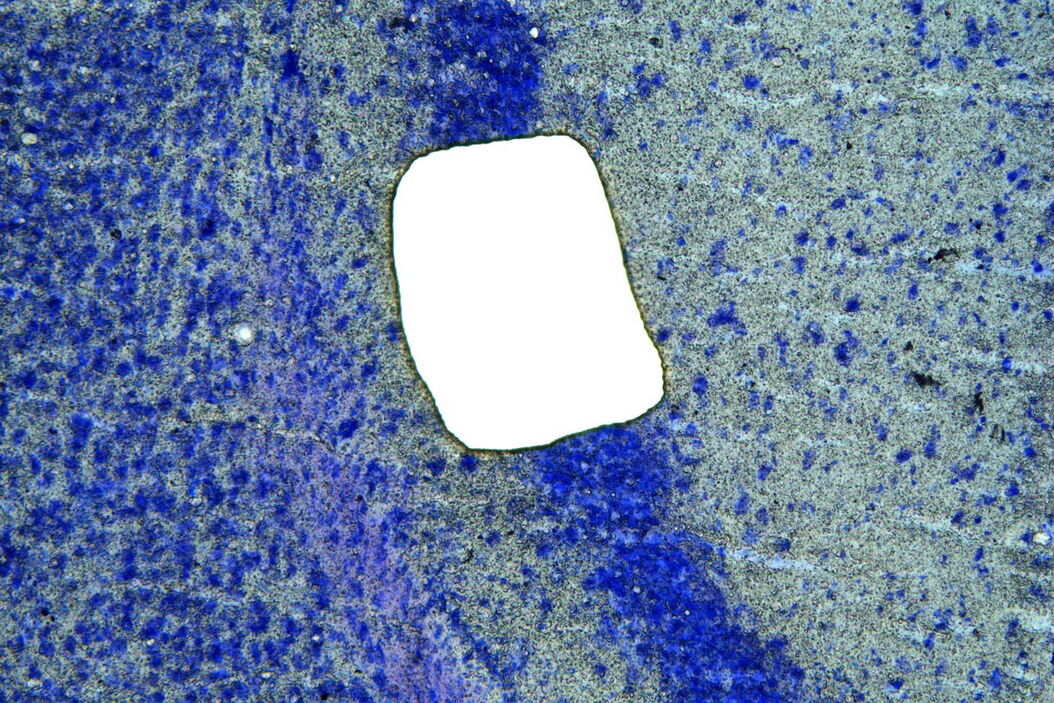
After fixing and staining, the applied tissue was isolated using the gentle and contamination-free technique of UV laser microdissection (UV LMD) [2]. This was done by marking the tissue and dividing it into rectangles using the 10x objective of the microscope before completely ablating it with the “Draw + Cut” mode using a laser beam of sufficient energy. The dissectates were collected in 0.5 ml RNase-free caps to allow purification of the RNA content. These were stored at –80° C pending RNA isolation.
Parallel RNA isolation
Both groups – the untreated samples stored at –80° C and the fixed, stained and laser-microdissected samples – were thawed in parallel and treated with the same reagents simultaneously. The RNeasy Mini Kit® of Qiagen was used for purification, following the manufacturer’s instructions. The purified RNA was eluted with 30-µl RNase-free water into a fresh RNase-free test tube.
Comparison of the RIN values regarding RNA quality
To compare the RNA quality of native cryo-tissue sections with that of CV-stained, EtOH-fixed, laser-microdissected tissue sections, RNA from each of the differently treated samples was applied to the same RNA nanochip (Agilent Bioanalyzer). The RNA nanochip separates RNA samples by capillary gel electrophoresis and grades the quality of the samples according to the way they move on the chip. As mentioned above, the quality is expressed as a RIN value where 10 indicates completely intact RNA and 1 completely degraded RNA.
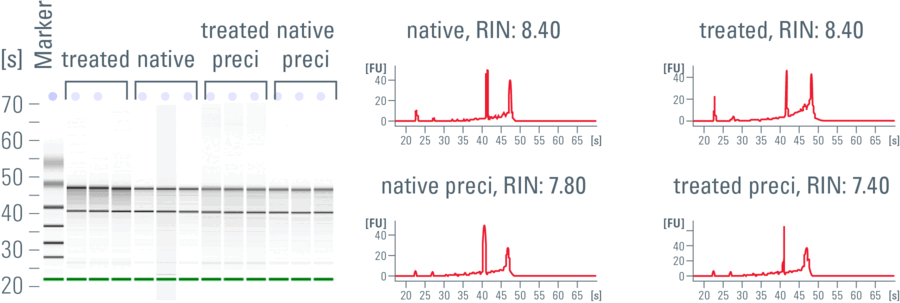
The RIN analysis conducted here (sample measurement in Fig. 2) showed that the RIN values of RNA from directly frozen native tissue were not much different from those treated according to the UV-LMD standard protocol.
RNA precipitation
The isolated RNA samples were then divided into 2 groups and half of each group was precipitated. After RNA precipitation, according to the instructions given in reference 3, the RIN values of RNA from native, precipitated cryo-tissue samples were slightly lower than the RIN values of RNA from CV-stained, EtOH-fixed, UV-LMD-treated tissue [3]. The difference can be attributed to the poly-I of the precipitation mixture – some of these short-strand RNA molecules are purified and detected with the other RNA during the RIN measurement in the chip and counted as part of the degraded proportion. In qPCR reactions, the poly-I does not show a negative influence. Therefore, fixation, staining, and UV laser microdissection of the brain sections with or without RNA precipitation do not impair RNA quality. Figure 2 shows a comparison of the RNA quality of native, untreated along with fixed and stained murine-brain sections.
Fig. 2: Comparison of the RNA quality of native, untreated as well as fixed and stained murine-brain sections: The example shows a virtual RNA gel (left) and the corresponding electropherogram (right) of an Agilent RNA chip using RNA isolated from native cryo-tissue (native) and CV-stained, EtOH-fixed, UV-LMD-treated tissue of the same murine brain, both before and after precipitation of the RNA (preci). The electropherograms result from the application of fluorescent units (FU) over time (in seconds [s]) [4,5].
Conclusions
Altogether, it was shown that UV laser microdissection with the method described [2] does not significantly impair the RNA quality of cryo-preserved brain tissue. Such a technique offers an appropriate method of sample preparation and non-destructive isolation of single cells to obtain RNA of consistent quality.