High‑pressure freezing (HPF) can help preserve hydrated cells and tissues close to their biological state at the moment of immobilization, supporting more reliable ultrastructural interpretation than conventional chemical preparation.
This eCollection brings together Bio‑protocol’s detailed, step‑by‑step HPF methods and applications across model systems.
What this eCollection contains (and how to use it)
This Bio protocol reprint collection focuses on advanced EM sample preparation across model systems—from yeast to plant and mammalian cells—with a protocol first format designed to support reproducibility and reliable execution in real labs.
If you are building or standardizing an HPF workflow, use the collection as a practical guide for:
- selecting an HPF route;
- moving samples through freeze substitution and embedding/sectioning;
- extending into electron tomography and 3D analysis when required.
Why high-pressure freezing matters for ultrastructure
HPF is positioned as a practical method to preserve hydrated cells and tissues in a state that closely reflects their biology at the moment of immobilization—helping address limitations of conventional chemical preparation and supporting more precise interpretation of ultrastructure.
As HPF became broadly adopted for EM sample preparation, it also became a foundation for reproducible workflows in ultrastructural and three dimensional imaging across many model systems.
The collection includes examples for challenging, fast-changing biology and complex 3D structures, where sample preservation and targeting decisions drive interpretation.
The collection spans multiple applications and readouts, including:
- Serial section electron tomography for fly mechanosensory organelles
- Serial‑section electron tomography and quantitative analysis of microtubule organization in 3D‑reconstructed mitotic spindles
- CLEM targeting of ultrastructural events at the graft interface of Arabidopsis thaliana, can support unambiguous targeting at the ultrastructural level and is compatible with immunolocalization and electron tomography for 3D views.
Additional protocols listed in the contents, including workflows for mammalian cells, C. elegans embryos, plasmodesmata, and yeast ultrastructure.
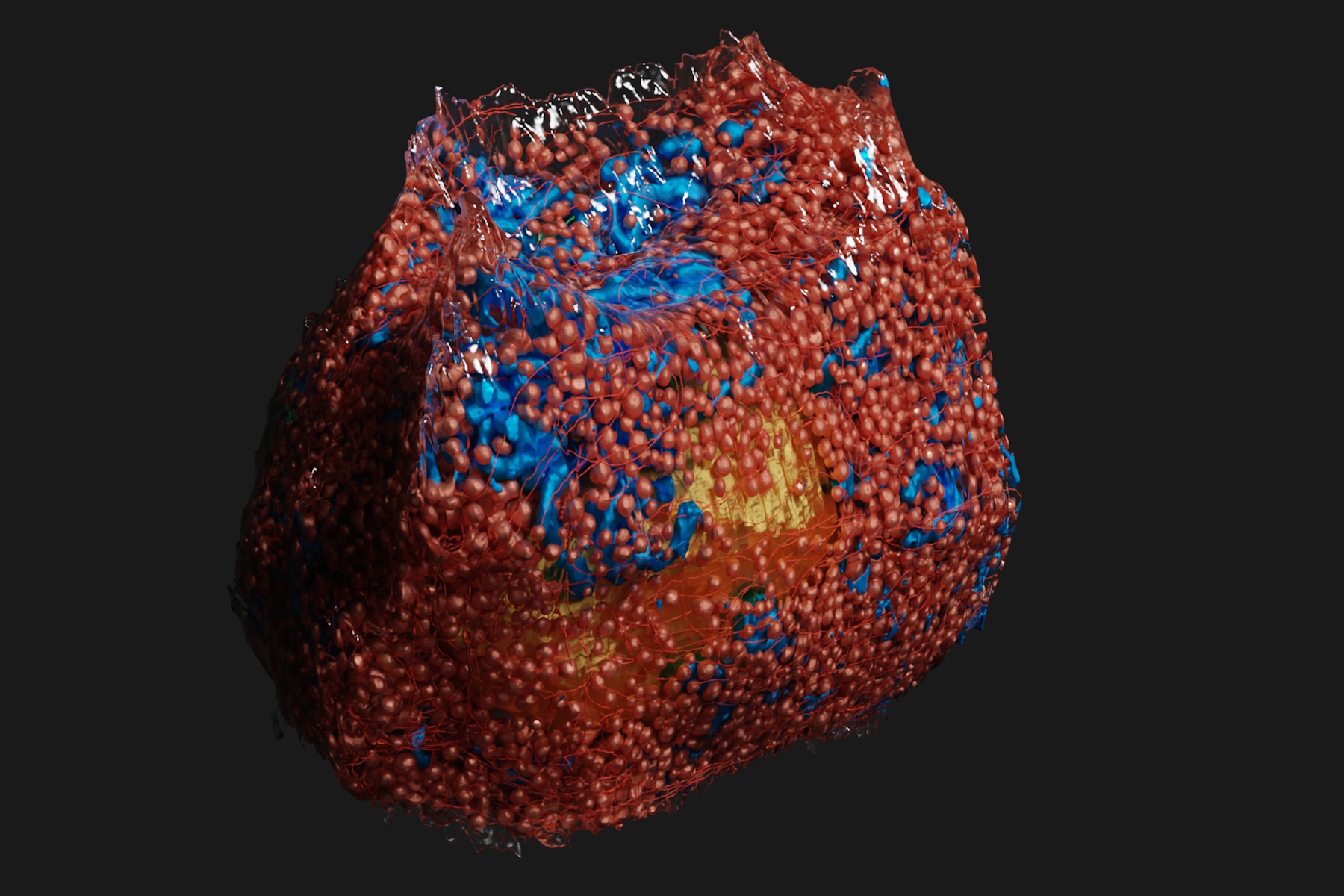
3D rendering of a mouse beta cell with mitochondria (blue), insulin SGs (orange), microtubules (red), nucleus (yellow), and plasma membrane (transparent). Image courtesy Andreas Müller, doi: https://doi.org/10.1083/jcb.202010039
Where the high-pressure freezer EM ICE fits
When your workflow depends on tight stimulation-to-freeze timing, the high-pressure freezer EM ICE helps reduce timing uncertainty by integrating stimulation control with rapid load-to-freeze performance.
Capture stimulation dependent ultrastructure with tighter timing control
- Built in light stimulation enables stimulation directly within the high-pressure freezer (no add on boxes or separate controllers).
- Software controlled stimulation streamlines setup by integrating control into the instrument software.
- Millisecond precision timing supports work where defining stimulation relative to freezing matters (light and electrical stimulation).
Reduce friction in the freezing step
- Alcohol free freezing runs without alcohol or added synchronization fluid.
- From loading to frozen in 1 second supports rapid transition from sample loading to freezing.
Use this eCollection as a practical starting point to standardize HPF workflows
This collection is positioned as a resource shaped by the community’s drive to standardize best practices—connecting sound methodology, dependable instrumentation, and application know how so that high quality preservation can be more accessible and consistent across laboratories and disciplines.
Because it focuses on detailed, step by step protocols, it is designed to help researchers and core facilities implement HPF workflows with an emphasis on reproducibility and reliability in day to day experimental work.